Abstract
Sex-linked inheritance is a stark exception to Mendel’s Laws of Heredity. Here we discuss how the evolution of heteromorphic sex chromosomes (mainly the Y) has been shaped by the intricacies of the meiotic programme. We propose that persistence of Y chromosomes in distantly related mammalian phylogroups can be explained in the context of pseudoautosomal region (PAR) size, meiotic pairing strategies, and the presence of Y-borne executioner genes that regulate meiotic sex chromosome inactivation. We hypothesise that variation in PAR size can be an important driver for the evolution of recombination frequencies genome wide, imposing constraints on Y fate. If small PAR size compromises XY segregation during male meiosis, the stress of producing aneuploid gametes could drive function away from the Y (i.e., a fragile Y). The Y chromosome can avoid fragility either by acquiring an achiasmatic meiotic XY pairing strategy to reduce aneuploid gamete production, or gain meiotic executioner protection (a persistent Y). Persistent Ys will then be under strong pressure to maintain high recombination rates in the PAR (and subsequently genome wide), as improper segregation has fatal consequences for germ cells. In the event that executioner protection is lost, the Y chromosome can be maintained in the population by either PAR rejuvenation (extension by addition of autosome material) or gaining achiasmatic meiotic pairing, the alternative is Y loss. Under this dynamic cyclic evolutionary scenario, understanding the meiotic programme in vertebrate and invertebrate species will be crucial to further understand the plasticity of the rise and fall of heteromorphic sex chromosomes.
Similar content being viewed by others
Sex chromosomes and recombination are exceptions to Mendelian inheritance
Mendel’s paramount observations can be considered one of the great discoveries in biology and are summarised into three laws of heredity: (i) Law of Independent Assortment; (ii) Law of Dominance; and (iii) Law of Segregation (Castle 1903). Under the first law (independent assortment), genes for different traits are inherited independently of one another. Under the second law (dominance), in crosses between homozygous parents for a contrasting character, only one character of the parent appears in the first generation. Finally, the third law (segregation) postulates that each egg or sperm cell receives just one of two copies of each chromosome during germ cell production, with the copy randomly allocated to gametes.
George Mendel himself soon realised that his observations in peas presented some limitations, and over the century and a half that followed his investigations exceptions to Mendelian inheritance have arisen. These include phenomena such as co-dominance, incomplete dominance, epistasis, pleiotropy and lethal alleles, among others (Castle 1903; Bateson 1909; Castle and Little 1910; Wright 1968; Stearns 2010). Sex-linked inheritance represents one of these exceptions, as genes carried on differentiated heteromorphic sex chromosomes show different inheritance patterns to those on autosomes (non-sex chromosomes). In addition, the sex-limited chromosome (the Y or W) does not undergo recombination, with the exception of the pseudoautosomal region (PAR) (Hinch et al. 2014; Raudsepp and Chowdhary 2015), so genes on them do not assort independently. This contrasts with autosomes, in which a recombination event occurs during the formation of germ cells (in a single generation) that can break gene assemblies that are physically linked.
Here we discuss how the intricacies of the meiotic programme have shaped the evolution of differentiated heteromorphic sex chromosomes (mainly the Y), resulting in exceptions to Mendelian inheritance. We walk through basic concepts of meiotic progression and chromosome dynamics in germ cells (meiocytes), coupled with a description of variation in recombination rates between phylogroups and the sexes (heterochiasmy). We then present an integrative hypothesis on how the cellular control of the meiotic programme and the mechanistic constraints of recombination can shape sex chromosome evolution. Although we focus mainly on mammals (i.e., system for which most information is available), our proposed model could apply to any differentiated XY or ZW sex chromosome system.
Meiosis and recombination
Recombination is essential for sexual reproduction due to its dual role in: (i) assembling new combinations of allelic variants that generate and maintain genotypic diversity; and (ii) establishing physical associations between homologous chromosomes to enable their faithful segregation during meiosis. From extensive work done in model organisms (i.e., yeast, fruit flies, nematodes, mice) and humans it is known that recombination occurs during the first meiotic prophase, which is organised into five stages: leptonema, zygonema, pachynema, diplonema and diakinesis (Fig. 1A). At leptonema, programmed double-strand breaks (DSBs) are generated and homologous chromosomes start to condense, pair and synapse. Chromosomes adhere to the nuclear envelope by their telomeres in the bouquet structure (reviewed in Reig-Viader et al. 2016), prompting the formation of the proteinaceous structure of the synaptonemal complex (SC) that, together with meiotic cohesins, acts as a scaffold for chromosome synapsis and recombination.
A Stages of spermatogenesis. Spermatogonia commit to meiosis and after DNA duplication, primary spermatocytes undergo meiotic prophase I: leptonema, zygonema, pachynema and diplonema. Double-strand breaks (DBSs) are formed at leptonema, repaired in zygonema, resulting in crossovers (COs) in pachynema. Two meiotic checkpoints are activated in this stage: (i) response to unrepaired DSBs; and (ii) MSUC (meiotic silencing of unsynapsed chromatin) and MSCI (meiotic sex chromosome inactivation). The first and second meiotic divisions result in secondary spermatocytes and round spermatids (RS), respectively. The spindle assembly checkpoint (SAC) is activated during division (metaphase-to-anaphase transition). Spermiogenesis includes the histone to protamine transition and the differentiation of RS into elongated spermatids (ES) and finally spermatozoa. Adapted from (Vara and Ruiz-Herrera 2022). Numbers between parentheses indicate the diploid (2n) haploid (n) number for each cell type and the number of chromatids per chromosome (4c, 2c, or c). B Schematic representation of chromosome pairing and MSCI dynamics during prophase I progression. Description of general patterns is based on evidence in mouse meiosis (adapted from Waters and Ruiz-Herrera 2020). C The formation and repair of DSBs occurs in the context of the SC (Adapted from Dapper and Payseur 2019). The proteins central to different steps during the formation and repair of DSBs, including homology search and strand invasion; synapsis; CO and NCO decision; and CO resolution. Briefly, the DSBs machinery is formed by the MCD recombinosome (MEI4-Containing DSB-promoting), a proteinaceous complex that includes MEI4, REC114 and IHO1 (Robert et al. 2016). Once the double-stranded DNA is resected, SPO11 is erased and posteriorly depleted from the recombination site, leaving 3’ single-stranded overhangs at both sites of DSBs (Lam and Keeney 2015). These 3’ single-stranded overhangs are coated by RPA (Robert et al. 2016), which recruits the recombinases RAD51 and DMC1 to facilitate the homologous search of single-stranded DNA filaments. After resected strands have undergone homology searching and strand invasion, DSBs will be repaired either as crossovers (COs) or non-crossovers (NCOs), with each process modulated by different proteins. COs are Holliday junctions (HdJ) resolved by the MutL complex (composed of MLH1 and MLH3), which is recruited by TEX11 (Dapper and Payseur 2019). HEI10 is also recruited to the SC together with the MutL complex, antagonistically regulating RNF212 (Reynolds et al. 2013).
The formation of DSBs activates the DNA damage response (DDR) mechanism (Baudat et al. 2010; Myers et al. 2010; Parvanov et al. 2010), an integral part of the meiosis programme. Both DSB formation and DDR are tightly regulated by meiotic checkpoints, including (i) the response to unrepaired DSBs; (ii) transcriptional repression called meiotic silencing of unsynapsed chromatin (MSUC); and (iii) the spindle assembly checkpoint (Subramanian and Hochwagen 2014) (Fig. 1A). Sex chromosomes are subjected to transcriptional silencing during the first male meiotic division by a phenomenon called meiotic sex chromosome inactivation (MSCI), a sex chromosome-specific extension of MSUC (Turner 2005) (Fig. 1B). This is observed in pachynema spermatocytes as the ‘sex body’, which is enriched for repressive histone modifications (Handel 2004).
Importantly, the SC establishes the chromosomal context in which synapsis and recombination between homologues take place (Fig. 1C). The successful progression of early prophase I is dependent on the assembly of chromatin loops into chromosomal axes and the formation and repair of DSBs (Keeney et al. 1997; Romanienko and Camerini-Otero 2000; Longhese et al. 2009). Crucially, the higher-order meiotic chromosome structure regulates the number and distribution of DSBs, and hence the final number of crossovers (COs) per cell (Zickler and Kleckner 1999; Kleckner 2006; Vara et al. 2021). This, in turn, results in a close interplay between SC length and DNA loop size, influencing CO distribution (Zickler and Kleckner 1999; Kleckner 2006). So it appears that SC axis length is a quantitative characteristic of synapsis that is strongly associated with recombination rate (Ruiz-Herrera et al. 2017; Wang et al. 2019). Importantly, in humans, the varied recombination rates within and between individuals have been linked to differences in SC length (Lynn et al. 2002). This was later confirmed in other taxa (Ruiz-Herrera et al. 2017; Wang et al. 2019), with a potential impact on the evolution of recombination rates (Wang et al. 2019; Sardell and Kirkpatrick 2020).
Patterns of variation in recombination rates
As meiotic recombination influences genome evolution, mammalian recombination landscapes are a reflection of the selective forces that affect the DNA sequence itself, the chromosomal distribution of COs (see below), and the three-dimensional genome folding in germ cells (Vara et al. 2021; Vara and Ruiz-Herrera 2022). Traditionally, theoretical work on the evolution of recombination rates has outnumbered the empirical evidence of recombination variation, especially in natural populations. This was mainly due to the intrinsic difficulties of directly measuring recombination events. However, this has improved over recent years as different approaches have been developed to estimate the number and genomic distribution of COs. These approaches can be classified as direct measurements (i.e., direct measure of recombination events or DSB sites in meiotic cells; Pan et al. 2011; Dumont and Payseur 2011; Smagulova et al. 2011; Brick et al. 2012; Segura et al. 2013; Fowler et al. 2014; Ruiz-Herrera et al. 2017) or indirect measurements of recombination (i.e., estimation of recombination rates using linkage data; Ellegren et al. 2012; Chan et al. 2012; Munch et al. 2014; Sparks et al. 2019; Turbek et al. 2021).
Empirical studies using both direct and indirect approaches have described variation in recombination rates within meiotic cells of the same individual, between individuals, populations, sexes and species, influencing patterns of heritability (Capilla et al. 2016; Stapley et al. 2017; Wang et al. 2019; Vara et al. 2021; Johnston et al. 2016; Kawakami et al. 2019). DSBs induced in early prophase I are higher in number than the final CO number, in some cases substantially more (≥10-fold) (Cole et al. 2012). The DSBs:COs ratio can vary between species, from 10:1 in mice to 3:1 in carnivores (Segura et al. 2013), likely influencing the observed differences in recombination rates.
Early genetic studies (Carpenter 1988) noticed that COs were non-randomly distributed along chromosomes (i.e., recombination hotspots), which were later identified by the immunodetection of MLH1 (Baker et al. 1996; Anderson et al. 1999; Lynn et al. 2002). After decades of study in different organisms it is currently accepted that the chromosomal distribution of COs exhibits four specific features, which are conserved in most species (Fig. 2A): (i) the presence of an obligatory CO; (ii) the phenomenon of CO interference; (iii) the centromeric effect; and (iv) CO homeostasis.
A Specific features that reduce local frequency (red box) of COs (red crosses). Chromosomal distribution of COs is influenced by the obligatory CO, centromeric effect and CO interference. B Schematic representation of general patterns of heterochiasmy in eutherian mammals with high recombination rates in females. C Chromosomes with short chromosomal axes (males) tend to show longer DNA loops than those with long chromosomal axes (females). These longer axes and shorter DNA loops in females result in increased COs (red stars). CO crossover, CEN centromere, TEL telomere, CO crossovers.
(i) Obligatory CO: there is normally a minimum of one CO per chromosome arm or chromosome, the so-called obligatory chiasma (Bishop and Zickler 2004; Zickler and Kleckner 2015). This serves to establish a necessary physical connection between homologous chromosomes during prophase I to avoid aneuploidies after chromosome segregation. This is a widely conserved pattern in eukaryotes (Zickler and Kleckner 1999; de Villena and Sapienza 2001; Segura et al. 2013).
(ii) CO interference: when a CO forms at one site of the chromosome this interferes with the establishment of COs at adjacent sites due to ‘interference’, resulting in evenly spaced COs (Muller 1916; Kleckner et al. 2003; Wang et al. 2015). This process is pervasive, and although the mechanisms behind it are not currently fully understood, two types of models to explain CO interference: mechanical models and diffusion-based models (reviewed in von Diezmann and Rog 2021).
(iii) Centromeric effect: centromeres normally act as ‘cold’ regions, with a reduction of COs (Beadle 1932; Mather 1939; Cappelletti et al. 2019). Low rates of recombination at centromeres were initially described in Drosophila using genetic maps, and later confirmed in different taxa ranging from plants to humans (Lynn et al. 2002; Colome-Tatche et al. 2012). The mechanisms governing centromere effect are far from understood, but this conserved phenomenon probably indicates the presence of strong selective constraints to avoid disruption of pericentric sister chromatid cohesion (Talbert and Henikoff 2010).
(iv) CO homeostasis: this phenomenon buffers the system against deficits (and excesses) of DSBs in meiocytes (Martini et al. 2006; Yokoo et al. 2012; Cole et al. 2012; Wang et al. 2015). It was initially defined as the maintenance of CO frequency even though precursors of DSBs are fewer (Martini et al. 2006). CO homeostasis balances the ratio between CO and NCOs to maintain the obligatory CO on each chromosome. This homeostasis is mechanistically linked to CO interference. As such, COs are maintained in a given cell at the expense of NCOs (Martini et al. 2006).
In addition to the four features described above, sex can also influence recombination rates. Heterochiasmy (sexual dimorphism in recombination rates) has been reported in many taxa, from invertebrates to mammals (Morgan 1912; Lynn et al. 2002). Early cytological work on human meiocytes observed a co-variation between SC length and recombination rates in both sexes (Wallace and Hultén 1985; Baker et al. 1996; Lynn et al. 2002) (Fig. 2B). SC length is longer and CO numbers are higher in oocytes than in spermatocytes (Wang et al. 2017), resulting in higher recombination rates in females (Fig. 2C). Increased SC length and heterochiasmy has also been observed in mouse (Lynn et al. 2002), zebrafish (Wallace and Wallace 2003), planarian (Jones and Croft 1989) and plants (Drouaud et al. 2007; Capilla-Pérez et al. 2021).
We have described that it is important to consider both the cellular context and the molecular constraints underlying the genomic distribution and frequency of meiotic recombination. But, in order to understand the interplay between sex, recombination and the meiotic programme, we need to explore Y chromosome evolution.
Theories of Y chromosome evolution
Although the current therian X and Y chromosomes have very different structure and gene content, they were once an ordinary pair of autosomes (Ohno 1967). In the therian ancestor (approximately 180 MYA), the proto-Y obtained the testis determining gene SRY (Foster et al. 1992). It has been hypothesised that once this new sex-determining system was established, male beneficial alleles accumulated nearby (in linkage disequilibrium) so that they were more likely inherited in males. Ultimately recombination was suppressed between the X and Y across this region, generating the first male-specific region of the Y (Rice 1996). This absence of recombination signalled the initial demise of Y-borne gene function, leading to its degradation.
The mechanisms leading to suppressed recombination have been extensively debated in the literature (Kratochvíl et al. 2021). It was long thought that inversions on the Y resulted in large regions of recombination suppression in single events (Lahn and Page 1999). However, other mechanisms for suppression of recombination have been proposed, including (i) pre-existing low recombination rates on autosomes that become sex chromosomes (Bergero et al. 2019; Rifkin et al. 2021; Xue et al. 2021); (ii) gradual expansion of suppressed recombination rather than large stepwise suppression (Darolti et al. 2020); (iii) different reproductive strategies (Mackiewicz et al. 2018); and (iv) even a neutral model of suppressed recombination (Jeffries et al. 2021). These models are not necessarily mutually exclusive, with different sex chromosome systems likely losing recombination via one or more of these mechanisms (Kratochvíl et al. 2021). Irrespective of how recombination was suppressed, in mammals Y degradation followed, resulting in the sex-specific chromosome becoming a relic of its former self.
Once thought to be a dominant element in the genome (because of its dominant testis determining action), the current degenerated nature of mammalian Y suggests that it is largely a wimpy relic of the X (the so-called wimpy Y—reviewed in Marshall Graves (2000)) (Table 1). Y chromosome decay has not been linear; instead, there were waves of gene loss, presumably from new male-specific regions of the Y soon after recombination was suppressed with the X (reviewed in Charlesworth 2021).
Y chromosomes have also been proposed to be fragile (Table 1) (Blackmon and Demuth 2014, 2015). Under the fragile Y hypothesis, a small PAR results in a less faithful pairing of the X and Y during male meiosis, subsequently increasing the stress of aneuploid gamete production as a result of improper segregation. This may impose a selective pressure to remove functions (i.e., genes) from the Y chromosome, predisposing it to being lost from the population. To prevent this aneuploidy stress and movement of function away from the Y, faithful achiasmatic mechanisms for XY segregation needs to evolve. In marsupial, a dense plate structure, rich in SC proteins, ensures faithful segregation in the absence of synapsis and recombination during the first meiotic division (Page et al. 2003; Marín-Gual et al. 2022). Alternatively, the PAR can be rejuvenated (extended) by the addition of autosomal material, as occurred in the eutherian ancestor under the addition-attrition hypothesis (Graves 1995). However, as new regions of the PAR stop recombining, PAR size is reduced and the Y degrades further, becoming fragile once more. PAR rejuvenation is only a temporary reprieve from fragility. So, in the absence of achiasmatic sex chromosome segregation, it appears that loss from the population is the inevitable fate of Y (or W) chromosomes in heteromorphic sex chromosome systems (Fig. 3).
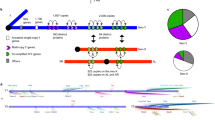
Initially, an ordinary pair of autosomes (purple chromosomes) became the therian sex chromosomes (blue chromosome). The proto-Y acquired a testis determining allele (new genetic sex determination [GSD], blue horizontal line) in the therian ancestor, around which male beneficial alleles accumulated (black horizontal lines). Recombination (grey circles between chromosome pairs) was suppressed across this region, signalling that the Y had become wimpy (purple ovals). As PAR size is reduced, the Y becomes fragile. Once a Y is fragile, there are four possible trajectories of evolution: (i) achiasmatic XY meiotic pairing (i.e., all marsupials); (ii) PAR rejuvenation (i.e., addition of pink autosomal material to the PAR in the ancestor to all eutherians); (iii) Y loss; or, (iv) gaining executioner protection (red horizontal line) to become persistent (i.e., still observed in almost all eutherians). Executioner protection could evolve on the Y at any stage that it is fragile; however, it is only shown after rejuvenation of the PAR in the eutherian ancestor. Persistent Y chromosomes, and those with achiasmatic meiotic pairing, are stable with no observation of being lost (green ovals). In a few unusual rodents (e.g., Lasiopodomys mandarinus, Dicrostonyx torquatus, Myopus schistocolor, Mus minutoides; Gil-Fernández et al. 2021; Saunders and Veyrunes 2021), persistent Ys have gained achiasmatic meiotic pairing. In a few other unusual rodent groups (e.g., Ellobius and Ryukyu spiny rats; Arakawa et al. 2002; Matveevsky et al. 2017) executioner protection has been removed, resulting once more in a wimpy and fragile Y, permitting its loss in some species (orange ovals). In these species, different autosomes must have become new sex chromosomes (either XY or ZW), completing the cycle. Green arrows represent evolutionary trajectories that have been observed in different mammal groups. Where there are two green arrows exiting a node, the groups of mammals that those trajectories have occurred in are shown. Grey arrows are proposed evolutionary trajectories despite a lack of observation in nature. Dashed arrows represent the completion of the circle, with a new pair of autosomes to become sex chromosomes.
However, Y chromosomes remain in all but a few therian mammal species (i.e., Ryukyu spiny rat and some Ellobius species) (Arakawa et al. 2002; Matveevsky et al. 2017; Furman et al. 2020; Vicoso 2019) since it arose approximately 181 MYA (Cortez et al. 2014). Indeed, the current human Y is relatively stable (Bellott et al. 2014, 2017; Cortez et al. 2014). This contrasts with observations in other vertebrate clades, in which there is regular sex chromosome turnover (Ezaz et al. 2006, 2010). In amphibians, there are some frogs with at least 13 sex chromosome turnover events in 28 species, which occurred within approximately 55 million years (Jeffries et al. 2018).
Persistent Ys in the meiotic context
Meiosis in general, and recombination (or the absence of it) more specifically, are both key elements to sex chromosome evolution. However, a unifying framework that accounts for how the meiotic cellular programme can influence sex chromosome evolution is still missing. This has been mainly due to the intrinsic difficulties of directly measuring recombination events and monitoring meiosis progression in germ cells of key representative phylogroups. So, despite being perceived as a fragile wimpy, why does the Y chromosome persist across large evolutionary time scales in the majority of mammalian species? Here we present an integrative view on how the meiotic programme and its mechanistic constraints can shape the evolution of differentiated sex chromosomes. We propose that the persistence of Y chromosomes in different phylogroups can be explained in the context of several interrelated aspects of sex chromosome biology and meiosis: (i) variation in PAR size; (ii) high recombination rates in the PAR to ensure proper segregation; and (iii) MSCI and the presence of Y-borne executioner genes.
Variation in PAR size: implications for Y evolution
In eutherian mammals, the PAR represents a small portion of the Y chromosome. PAR rejuvenation occurred in the eutherian ancestor (after marsupial divergence, but pre-eutherian radiation) under the addition-attrition model of sex chromosome evolution (Graves 1995). After PAR extension by this autosomal addition (Fig. 3), recombination was further suppressed between the X and Y. Different rates of Y degradation and recombination suppression within phylogroups resulted in variation of PAR size among mammals, from ≈700 Kbp in mice to ≈10 Mbp in alpaca (Raudsepp et al. 2012). In some eutherian clades (e.g., some bat, rodents, bovids and primates) the PAR has been rejuvenated, (Murata et al. 2012; Britton-Davidian et al. 2012; Rahn et al. 2016; Vozdova et al. 2016) presumably resulting in new Y specific material after the suppression of recombination between the X and Y in the extended PAR.
Variation in PAR size is an important driver for Y evolution under the fragile Y hypothesis (Blackmon and Demuth 2014, 2015) (Table 1). Y losses in the offspring (i.e., XO individuals) are frequently found in species with small PARs (0.7–2.7 Mb, i.e., human, mouse and horse; reviewed in Raudsepp and Chowdhary 2016) with negative (often fatal) consequences for the individual if not mosaic as reported in human and horse (Raudsepp et al. 2012). This aneuploidy stress would put pressure on the critical functions to be moved from the Y to the autosomes, as has been observed in mammals (Hughes et al. 2015).
In contrast, in domestic species with larger PARs (>5 Mb, i.e., ruminants, pigs and carnivores; reviewed in Raudsepp and Chowdhary 2016) XO individuals are infrequently observed, potentially as a result of more haploinsufficient genes being lethal (Raudsepp et al. 2012). However, it is tempting to speculate that under the fragile Y hypothesis, a larger PAR results in more faithful pairing and segregation of the sex chromosomes, so the generation of Y-less gametes is less frequent. Or perhaps a combination of both results in the observation of fewer XO individuals. That is, larger PARs result in less mis-segregation and a more lethal haploinsufficiency effect.
Recombination in the PAR
Progressive degeneration of the Y chromosome has led to the evolution of meiotic mechanisms that ensure faithful XY segregation during male meiosis. In eutherian mammals, most species have a PAR (reviewed in Raudsepp and Chowdhary 2016) that has evolved at high rates of recombination to ensure an obligate CO during male meiosis (Kauppi et al. 2011). In contrast to autosomes where DSBs occur every 10 Mbp (Kauppi et al. 2011), PARs are characterised by a recombination rate 10–20-fold higher than the genome average (e.g., 1–2 DSBs every 1 Mbp in rodents) (Kauppi et al. 2011). Initial studies proposed that this prevalence of DSB formation in the PAR was induced by a high-order chromosome structure distinct from that of the autosomes, such as the presence of abundant short DNA loops in a relatively long chromosomal axis (Kauppi et al. 2011). This was later demonstrated by the accumulation of cis- and trans-acting DSB-promoting proteins in the PAR during early prophase I (Papanikos et al. 2019; Acquaviva et al. 2020). Whether this observation in mouse holds for other eutherian mammals remains to be experimentally demonstrated. However, evidence of a correlation between DSBs, DNA loop size and chromosomal axes length in different taxa (Ruiz-Herrera et al. 2017; Wang et al. 2019) suggests this to be the case.
We propose that in species where the heterogametic sex is achiasmate (that is, sex chromosomes do not pair and recombine), the absence of a PAR would not impose selective pressure to initiate excessive numbers of DSBs genome wide to ensure a CO in a small PAR. Marsupials, which lack a PAR, are characterised by some of the lowest recombination rates within mammals (Zenger et al. 2002; Segura et al. 2013) and low number diploid numbers (which range from 2n = 10 to 2n = 32, Deakin and O’Neill 2020). These low levels of recombination in marsupials could result from a reduction of DSB formation genome wide (Marín-Gual et al. 2022). As a result of no PAR, DSBs are only required at a frequency sufficient to result in obligatory COs on the few large autosomes.
MSCI and the presence of Y-borne executioner genes
Whether the heterogametic sex is achiasmate (i.e., marsupials and some rodent species), or not (i.e., eutherian mammals), MSCI occurs and is a necessary checkpoint to ensure proper meiotic progression. It was recently proposed that the unique features of MSCI might explain the persistence of the Y chromosome in most eutherian species (Waters and Ruiz-Herrera 2020). That is, Y-borne meiotic executioner genes, integral to meiotic checkpoints, lead to the persistence of Y chromosomes in species with MSCI (i.e., a persistent Y, Table 1). Meiotic executioners are pachytene lethal if they escape MSCI (Royo et al. 2010; Vernet et al. 2016), resulting in the production of inviable gametes. In addition to executing the cell when ectopically expressed, to be a truly selfish element that protects the Y, executioners are also required for the initiation of MSCI (Royo et al. 2010; Vernet et al. 2016). The result is a gene that has evolved to become its own sensor.
Under the persistent Y hypothesis (Waters and Ruiz-Herrera 2020), once a Y chromosome is under execution protection, it cannot be lost from the population. The Y as a unit is required for successful MSCI, to which it must then be subject. Under no circumstance can viable Y-less gametes be produced. Should an executioner be translocated to an autosome, MSCI proceeds but the executioner escapes silencing and the cell dies. The only location in the genome to which an executioner gene can move is the X chromosome. From there it must retain its function to properly initiate MSCI, so that it is appropriately silenced, permitting meiosis to proceed. Indeed, the movement of executioner genes from the Y to the X preceded Y chromosome loss in Y-less rodent species (reviewed in Waters and Ruiz-Herrera 2020). In eutherian mammals, Zfy acts as the executioner, and its translocation to an autosome explains complete azoospermia reported in a horse (Ruiz et al. 2019; Bugno-Poniewierska and Raudsepp 2021). It is important to note that persistent Ys are only possible in clades with MSCI already established, providing a haven from which genes can evolve a selfish executioner function.
The evolution of executioner function is a fascinating question, especially in light of the fact that Zfy was part of an addition to the sex chromosomes in the eutherian ancestor, and remains autosomal in marsupials (Waters et al. 2001). Being a pachytene lethal executioner gene, it is no surprise that Zfy expression is low in the spermatocytes of representative eutherian species (Murat et al. 2021). However, what function could it play from an autosome in non-eutherian species?
In opossum, platypus and chicken, Zfy is annotated as its X homologue, Zfx. In each of these species, Zfx expression is maintained in spermatocytes (Murat et al. 2021), suggesting that the Zfx/y precursor was not pachytene lethal. Therefore, it could only have gained this aspect of executioner function after arriving on the eutherian Y chromosome. It is unclear if the Zfx/y precursor already had a function in initiating MSCI before its translocation to the Y; however, it is tempting to speculate that it did. In opossum and platypus, in which MSCI occurs, Zfx expression is elevated in spermatocytes compared to other cell types (Murat et al. 2021).
In contrast, Zfx expression in chickens is only elevated in pre-meiotic cells (i.e., spermatogonia). But since birds have a ZW sex chromosome system and males are the homogametic sex, there is no requirement for MSCI in the testis. Because MSCI is a prerequisite for the evolution of executioner genes, further studies on the germline of the heterogametic sex in distantly related vertebrate linages will provide important insight into the potential for the evolution of executioner function.
Closing the circle
Here we have described how sex chromosomes result in exceptions to Mendelian inheritance, highlighting the importance of functional and mechanistic meiotic constraints on sex chromosome evolution. Under a dynamic (and possibly cyclical) evolutionary trajectory (Fig. 3) we propose that Y persistence can be explained in the context of recombination rates, PAR size and Y-borne meiotic executioner genes that regulate MSCI.
Due to suppressed recombination, the PAR represents a small portion of the X and Y. We propose that these fragile Ys with small PARs result in an increase of DSBs (and hence recombination rates) to ensure a CO event in the PAR, and reduce the deleterious effect of aneuploidy. If PAR size is reduced so that the Y becomes too fragile and aneuploidy stress causes loss of Y function, the Y chromosome can become stable by (i) temporary PAR rejuvenation, (ii) evolving achiasmatic XY segregation, or (iii) acquiring executioner protection (Fig. 3). A persistent Y will remain under strong constraints to maintain high recombination rates in the PAR to ensure an obligatory CO, as losing the Y will have fatal consequences for germ cells (i.e., aberrant MSCI). Almost all eutherian Y chromosomes are at this stage. In the rare event that executioner protection is lost (i.e., translocation away from the Y chromosome to the X), the Y chromosome can be maintained in the population by either PAR rejuvenation or evolving achiasmatic XY segregation, the alternative is Y loss.
Under this dynamic cyclic evolutionary scenario, the intricacies of the meiotic programme influence the fate of Y chromosomes. Understanding the progression and regulation of the meiotic programme in vertebrate and invertebrate species will be important to further decipher the plasticity of the rise and fall of heteromorphic sex chromosomes.
References
Acquaviva L, Boekhout M, Karasu ME, Brick K, Pratto F, Li T et al. (2020) Ensuring meiotic DNA break formation in the mouse pseudoautosomal region. Nature 582:426–431
Aitken RJ, Marshall Graves JA (2002) Human spermatozoa: the future of sex. Nature 415:963–963
Anderson LK, Reeves A, Webb LM, Ashley T (1999) Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151:1569–1579
Arakawa Y, Nishida-Umehara C, Matsuda Y, Sutou S, Suzuki H (2002) X-chromosomal localization of mammalian Y-linked genes in two XO species of the Ryukyu spiny rat. Cytogenetic Genome Res 99:303–309
Baker SM, Plug AW, Prolla TA, Bronner CE, Harris AC, Yao X et al. (1996) Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over. Nat Genet 13:336–342
Bateson W (1909) Mendel’s principles of heredity. University Press, Cambridge
Baudat F, Buard J, Grey C, Fledel-Alon A, Ober C, Przeworski M et al. (2010) PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science 327:836–840
Beadle GW (1932) A possible influence of the spindle fibre on crossing-over in Drosophila. Proc Natl Acad Sci USA 18:160–165
Bellott DW, Hughes JF, Skaletsky H, Brown LG, Pyntikova T, Cho TJ et al. (2014) Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508:494–499
Bellott DW, Skaletsky H, Cho TJ, Brown L, Locke D, Chen N et al. (2017) Avian W and mammalian Y chromosomes convergently retained dosage-sensitive regulators. Nat Genet 49:387–394
Bergero R, Gardner J, Bader B, Yong L, Charlesworth D (2019) Exaggerated heterochiasmy in a fish with sex-linked male coloration polymorphisms. Proc Natl Acad Sci USA 116:6924–6931
Bishop DK, Zickler D (2004) Early decision; meiotic crossover interference prior to stable strand exchange and synapsis. Cell 117:9–15
Blackmon H, Demuth JP (2014) Estimating tempo and mode of Y chromosome turnover: explaining Y chromosome loss with the fragile Y hypothesis. Genetics 197:561–572
Blackmon H, Demuth JP (2015) The fragile Y hypothesis: Y chromosome aneuploidy as a selective pressure in sex chromosome and meiotic mechanism evolution. BioEssays 37:942–950
Brick K, Smagulova F, Khil P, Camerini-Otero RD, Petukhova GV (2012) Genetic recombination is directed away from functional genomic elements in mice. Nature 485:642–645
Britton-Davidian J, Robinson TJ, Veyrunes F (2012) Systematics and evolution of the African pygmy mice, subgenus Nannomys: a review. Acta Oecologica 42:41–49
Bugno-Poniewierska M, Raudsepp T (2021) Horse clinical cytogenetics: recurrent themes and novel findings. Animals 11:1–26.
Capilla L, Garcia Caldés M, Ruiz-Herrera A (2016) Mammalian meiotic recombination: a toolbox for genome evolution. Cytogenetic Genome Res 150:1–16
Capilla-Pérez L, Durand S, Hurel A, Lian Q, Chambon A, Taochy C et al. (2021) The synaptonemal complex imposes crossover interference and heterochiasmy in Arabidopsis. Proc Natl Acad Sci USA 118:e2023613118
Cappelletti E, Piras FM, Badiale C, Bambi M, Santagostino M, Vara C et al. (2019) CENP-A binding domains and recombination patterns in horse spermatocytes. Sci Rep. 9:15800
Carpenter ATC (1988) Thoughts on recombination nodules, meiotic recombination, and chiasmata. In: Kucherlapati R, Smith GR (eds) Genetic recombination. American Society of Microbiology, Washington, DC, p 529–548
Castle WE (1903) Mendel’s Law of Heredity. Science 18:396–406
Castle WE, Little CC (1910) On a modified mendelian ratio among yellow mice. Science 32:868–870
Chan AH, Jenkins PA, Song YS (2012) Genome-wide fine-scale recombination rate variation in Drosophila melanogaster. PLoS Genet 8(12):e1003090
Charlesworth D (2021) The timing of genetic degeneration of sex chromosomes. Philos Trans R Soc B: Biol Sci 376:20200093
Cole F, Kauppi L, Lange J, Roig I, Wang R, Keeney S et al. (2012) Homeostatic control of recombination is implemented progressively in mouse meiosis. Nat Cell Biol 14(4):424–430
Colome-Tatche M, Cortijo S, Wardenaar R, Morgado L, Lahouze B, Sarazin A et al. (2012) Features of the Arabidopsis recombination landscape resulting from the combined loss of sequence variation and DNA methylation. Proc Natl Acad Sci USA 109:16240–16245
Cortez D, Marin R, Toledo-Flores D, Froidevaux L, Liechti A, Waters PD et al. (2014) Origins and functional evolution of Y chromosomes across mammals. Nature 508:488–493
Dapper AL, Payseur BA (2019) Molecular evolution of the meiotic recombination pathway in mammals. Evolution 73:2368–2389
Darolti I, Wright AE, Mank JE (2020) Guppy Y chromosome integrity maintained by incomplete recombination suppression. Genome Biol Evolution 12:965–977
de Villena FP (2001) Female meiosis drives karyotypic evolution in mammals. Genetics 159:1179–1189
Deakin JE, O’Neill RJ (2020) Evolution of marsupial genomes. Annu Rev Anim Biosci 8:25–45
Drouaud J, Mercier R, Chelysheva L, Bérard A, Falque M, Martin O et al. (2007) Sex-specific crossover distributions and variations in interference level along Arabidopsis thaliana chromosome 4. PLoS Genet 3:e106
Dumont BL, Payseur BA (2011) Evolution of the genomic recombination rate in Murid rodents. Genetics 187(3):643–57
Ellegren H, Smeds L, Burri R, Olason PI, Backström N, Kawakami T et al. (2012) The genomic landscape of species divergence in Ficedula flycatchers. Nature 491(7426):756–760
Ezaz T, Sarre SD, O’Meally D, Marshall Graves JA, Georges A (2010) Sex chromosome evolution in lizards: Independent origins and rapid transitions. Cytogenetic Genome Res 127:249–260
Ezaz T, Stiglec R, Veyrunes F, Graves JAM (2006) Relationships between vertebrate ZW and XY sex chromosome systems. Curr Biol 16:R736–R743
Foster JW, Brennan FE, Hampikian GK, Goodfellow PN, Sinclair AH, Lovell-Badge R et al. (1992) Evolution of sex determination and the Y chromosome: SRY-related sequences in marsupials. Nature 359:531–533
Fowler KR, Sasaki M, Milman N, Keeney S, Smith GR (2014) Evolutionarily diverse determinants of meiotic DNA break and recombination landscapes across the genome. Genome Res 24:1650–1664
Furman M, Darolti W, Sandkam Almeida S, Mank J (2020). Sex Chromosome Evolution: SoMany Exceptions to the Rules. Genome Biology and Evolution 12:750–763.
Gil-Fernández A, Ribagorda M, Martín-Ruiz M, López-Jiménez P, Laguna T, Gómez R et al. (2021) Meiotic behavior of achiasmate sex chromosomes in the african pygmy Mouse Mus mattheyi offers new insights into the evolution of sex chromosome pairing and segregation in mammals. Genes 12:1434
Graves JAM (1995) The origin and function of the mammalian Y chromosome and Y‐borne genes – an evolving understanding. BioEssays 17:311–320
Handel MA (2004) The XY body: a specialized meiotic chromatin domain. Exp Cell Res 296:57–63
Hinch AG, Altemose N, Noor N, Donnelly P, Myers SR (2014) Recombination in the human pseudoautosomal region PAR1. PLoS Genet 10:e1004503
Hughes JF, Skaletsky H, Koutseva N, Pyntikova T, Page DC (2015) Sex chromosome-to-autosome transposition events counter Y-chromosome gene loss in mammals. Genome Biol 16:1–9
Jeffries DL, Gerchen JF, Scharmann M, Pannell JR (2021) A neutral model for the loss of recombination on sex chromosomes. Philos Trans R Soc B 376:20200096
Jeffries DL, Lavanchy G, Sermier R, Sredl MJ, Miura I, Borzée A et al. (2018) A rapid rate of sex-chromosome turnover and non-random transitions in true frogs. Nat Commun 9:1–11
Johnston SE, Bérénos C, Slate J, Pemberton JM (2016). Conserved Genetic Architecture Underlying Individual Recombination Rate Variation in a Wild Population of Soay Shee p (Ovis aries). Genetics, 203:583–598.
Jones GH, Croft JA (1989) Chromosome pairing and chiasma formation in spermatocytes and oocytes of Dendrocoelum lactem (Turbellaria, Tricladida); a cytogenetical and ultrastructural study. Heredity 63:97–106
Kauppi L, Barchi M, Baudat F, Romanienko PJ, Keeney S, Jasin M (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331:916–920
Kawakami T, Wallberg A, Olsson A, Wintermantel, D, de Miranda, JR, Allsopp M, et al. (2019). Substantial Heritable Variation in Recombination Rate on Multiple Scales in Honeybees and Bumblebees. Genetics, 212:1101–1119.
Keeney S, Giroux NC, Kleckner N (1997) Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell, 88:375–384.
Kleckner N (2006) Chiasma formation: chromatin/axis interplay and the role(s) of the synaptonemal complex. Chromosoma 115:175–194
Kleckner N, Storlazzi A, Zickler D (2003) Coordinate variation in meiotic pachytene SC length and total crossover/chiasma frequency under conditions of constant DNA length. Trends Genet 19:623–628
Kratochvíl L, Stöck M, Rovatsos M, Bullejos M, Herpin A, Jeffries DL et al. (2021) Expanding the classical paradigm: what we have learnt from vertebrates about sex chromosome evolution. Philos Trans R Soc B 376:20200097
Lahn BT, Page DC (1999) Four evolutionary strata on the human X chromosome. Science 286:964–967
Lam I, Keeney S (2015) Mechanism and regulation of meiotic recombination initiation. Cold Spring Harb Perspect Biol 7:a016634
Longhese MP, Bonetti D, Guerini I, Manfrini, N, Clerici M (2009). DNA double-strand breaks in meiosis: Checking their formation, processing and repair. DNA Repair, 8:1127–1138.
Lynn A, Koehler KE, Judis L, Chan ER, Cherry JP, Schwartz S et al. (2002) Covariation of synaptonemal complex length and mammalian meiotic exchange rates. Science 296:2222–2225
Mackiewicz D, Posacki P, Burdukiewicz M, Błażej P (2018) Role of recombination and faithfulness to partner in sex chromosome degeneration. Sci Rep 8:1–15
Marín-Gual L, González-Rodelas L, Pujol G, Vara C, Martín-Ruiz M, Berríos S et al. (2022) Strategies for meiotic sex chromosome dynamics and telomeric elongation in Marsupials. PLOS Genet 18:e1010040
Marshall Graves JA (2000) Human Y chromosome, sex determination, and spermatogenesis—a feminist view. Biol Reprod 63:667–676
Martini E, Diaz RL, Hunter N, Keeney S (2006) Crossover homeostasis in yeast meiosis. Cell 126:285–295
Mather K (1939) Crossing over and heterochromatin in the X chromosome of Drosophila melanogaster. Genetics 24:413–435
Matveevsky S, Kolomiets O, Bogdanov A, Hakhverdyan M, Bakloushinskaya I (2017) Chromosomal evolution in mole Voles ellobius (Cricetidae, rodentia): bizarre sex chromosomes, variable autosomes and meiosis. Genes 8:306
Morgan T (1912) Complete linkage in the second chromosome of the male of Drosophila. Science 36:719–720
Muller HJ (1916) The mechanism of crossing-over. Am Naturalist 592:284–305
Munch K, Schierup MH, Mailund T (2014) Unraveling recombination rate evolution using ancestral recombination maps. BioEssays 36(9):892–900
Murat F, Mbengue N, Winge SB, Trefzer T, Leushkin E, Sepp M, et al. (2021) The molecular evolution of spermatogenesis across mammals. Preprint at bioRxiv 2021.11.08.467712
Murata C, Yamada F, Kawauchi N, Matsuda Y, Kuroiwa A (2012) The y chromosome of the Okinawa spiny rat, Tokudaia muenninki, was rescued through fusion with an autosome. Chromosome Res 20:111–125
Myers S, Bowden R, Tumian A, Bontrop RE, Freeman C, MacFie TS et al. (2010) Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science 327:876–879
Ohno S (1967) Sex chromosomes and sex-linked genes. Springer, Berlin, Germany
Page J, Berríos S, Rufas JS, Parra MT, Suja JA, Heyting C et al. (2003) The pairing of X and Y chromosomes during meiotic prophase in the marsupial species Thylamys elegans is maintained by a dense plate developed from their axial elements. J Cell Sci 116:551–560
Pan J, Sasaki M, Kniewel R, Murakami H, Blitzblau HG, Tischfield SE et al. (2011) A hierarchical combination of factors shapes the genome-wide topography of yeast meiotic recombination initiation. Cell 144:719–731
Papanikos F, Clément JAJ, Testa E, Ravindranathan R, Grey C, Dereli I et al. (2019) Mouse ANKRD31 regulates spatiotemporal patterning of meiotic recombination initiation and ensures recombination between X and Y Sex chromosomes. Mol Cell 74:1069–1085.e11
Parvanov ED, Petkov PM, Paigen K (2010) Prdm9 controls activation of mammalian recombination hotspots. Science 327:835
Rahn MI, Noronha RC, Nagamachi CY, Pieczarka JC, Solari AJ, Sciurano RB (2016) Protein markers of synaptic behavior and chromatin remodeling of the neo-XY body in phyllostomid bats. Chromosoma 125:701–708
Raudsepp T, Chowdhary BP (2015) The eutherian pseudoautosomal region. Cytogenetic Genome Res 147:81–94
Raudsepp T, Chowdhary BP (2016) Chromosome aberrations and fertility disorders in domestic animals. Annu Rev Anim Biosci 4:15–43
Raudsepp T, Das PJ, Avila F, Chowdhary BP (2012) The pseudoautosomal region and sex chromosome aneuploidies in domestic species. Sex Dev 6:72–83
Reig-Viader R, Garcia-Caldés M, Ruiz-Herrera A (2016) Telomere homeostasis in mammalian germ cells: a review. Chromosoma 125:337–351
Reynolds A, Qiao H, Yang Y, Chen JK, Jackson N, Biswas K et al. (2013) RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis. Nat Genet 45:269–278
Rice WR (1996) Evolution of the Y sex chromosome in animals Y chromosomes evolve through the degeneration of autosomes. BioScience 46:331–343
Rifkin JL, Beaudry FEG, Humphries Z, Choudhury BI, Barrett SCH, Wright SI (2021) Widespread recombination suppression facilitates plant sex chromosome evolution. Mol Biol Evolution 38:1018–1030
Robert T, Nore A, Brun C, Maffre C, Crimi B, Guichard V et al. (2016) The TopoVIB-Like protein family is required for meiotic DNA double-strand break formation. Science 351:943–949
Romanienko PJ, Camerini-Otero RD (2000). The Mouse Spo11 Gene Is Required for Meiotic Chromosome Synapsis. Molecular Cell, 6:975–987.
Royo H, Polikiewicz G, Mahadevaiah SK, Prosser H, Mitchell M, Bradley A et al. (2010) Evidence that meiotic sex chromosome inactivation is essential for male fertility. Curr Biol 20:2117–2123
Ruiz AJ, Castaneda C, Raudsepp T, Tibary A (2019) Azoospermia and Y chromosome–autosome translocation in a Friesian stallion. J Equine Vet Sci 82:102781
Ruiz-Herrera A, Vozdova M, Fernández J, Sebestova H, Capilla L, Frohlich J et al. (2017) Recombination correlates with synaptonemal complex length and chromatin loop size in bovids—insights into mammalian meiotic chromosomal organization. Chromosoma 126:615–631
Sardell JM, Kirkpatrick M (2020) Sex differences in the recombination landscape. Am Naturalist 195:361–379
Saunders PA, Veyrunes F (2021) Unusual mammalian sex determination systems: a cabinet of curiosities. Genes 12:1770
Segura J, Ferretti L, Ramos-Onsins S, Capilla L, Farré M, Reis F et al. (2013) Evolution of recombination in eutherian mammals: Insights into mechanisms that affect recombination rates and crossover interference. Proc R Soc B: Biol Sci 280:20131945
Smagulova F, Gregoretti IV, Brick K, Khil P, Camerini-Otero RD, Petukhova GV (2011) Genome-wide analysis reveals novel molecular features of mouse recombination hotspots. Nature 472:375–378
Sparks AM, Watt K, Sinclair R, Pilkington JG, Pemberton JM, McNeilly TN et al. (2019) The genetic architecture of helminth-specific immune responses in a wild population of Soay sheep (Ovis aries). PLoS Genetics 15:e1008461
Stapley J, Feulner PGD, Johnston SE, Santure AW, Smadja CM (2017) Variation in recombination frequency and distribution across eukaryotes: patterns and processes. Philos Trans R Soc B: Biol Sci 372:20160455
Stearns FW (2010) One hundred years of pleiotropy: a retrospective. Genetics 186:767–773
Subramanian VV, Hochwagen A (2014) The meiotic checkpoint network: step-by-step through meiotic prophase. Cold Spring Harb Perspect Biol 6:a016675
Talbert PB, Henikoff S (2010) Centromeres convert but don’t cross. PLOS Biol 8:e1000326
Turbek SP, Semenov GA, Enbody ED, Campagna L, Taylor SA (2021) Variable signatures of selection despite conserved recombination landscapes early in speciation. J Heredity 112:485–496
Turner JMA (2005) Sex chromosomes make their mark. Chromosoma 114:300–306
Vara C, Paytuví-Gallart A, Cuartero Y, Álvarez-González L, Marín-Gual L, Garcia F et al. (2021) The impact of chromosomal fusions on 3D genome folding and recombination in the germ line. Nat Commun 12:1–17
Vara C, Ruiz-Herrera A (2022) Unpacking chromatin remodelling in germ cells: implications for development and evolution. Trends Genet 38:422–425
Vernet N, Mahadevaiah SK, de Rooij DG, Burgoyne PS, Ellis PJI (2016) Zfy genes are required for efficient meiotic sex chromosome inactivation (MSCI) in spermatocytes. Hum Mol Genet 25:5300–5310
Vicoso B (2019). Molecular and evolutionary dynamics of animal sex-chromosome turnover. Nature Ecology and Evolution. 3:1632–1641.
von Diezmann L, Rog O (2021) Let’s get physical – mechanisms of crossover interference. J Cell Sci 134:jcs255745
Vozdova M, Ruiz-Herrera A, Fernandez J, Cernohorska H, Frohlich J, Sebestova H et al. (2016) Meiotic behaviour of evolutionary sex-autosome translocations in Bovidae. Chromosome Res 24:325–338
Wallace BMN, Hultén MA (1985) Meiotic chromosome pairing in the normal human female. Ann Hum Genet 49:215–226
Wallace BMN, Wallace H (2003) Synaptonemal complex karyotype of zebrafish. Heredity 90:136–140
Wang S, Hassold T, Hunt P, White MA, Zickler D, Kleckner N et al. (2017) Inefficient crossover maturation underlies elevated aneuploidy in human female meiosis. Cell 168:977–989.e17
Wang S, Liu Y, Shang Y, Zhai B, Yang X, Kleckner N et al. (2019) Crossover interference, crossover maturation, and human aneuploidy. BioEssays 41:e1800221
Wang S, Zickler D, Kleckner N, Zhang L (2015) Meiotic crossover patterns: obligatory crossover, interference and homeostasis in a single process. Cell Cycle 14:305–314
Waters PD, Duffy B, Frost CJ, Delbridge ML, Graves JAM (2001) The human Y chromosome derives largely from a single autosomal region added to the sex chromosomes 80-130 million years ago. Cytogenetics Cell Genet 92:74–79
Waters PD, Ruiz-Herrera A (2020) Meiotic executioner genes protect the Y from extinction. Trends Genet 36:728–738
Wright S (1968) Genetic and biometric foundations. University of Chicago Press, Chicago
Xue L, Gao Y, Wu M, Tian T, Fan H, Huang Y et al. (2021) Telomere-to-telomere assembly of a fish Y chromosome reveals the origin of a young sex chromosome pair. Genome Biol 22:1–20
Yokoo R, Zawadzki KA, Nabeshima K, Drake M, Arur S, Villeneuve AM (2012) COSA-1 reveals robust homeostasis and separable licensing and reinforcement steps governing meiotic crossovers. Cell 149:75–87
Zenger KR, McKenzie LM, Cooper DW (2002) The first comprehensive genetic linkage map of a marsupial: the tammar wallaby (Macropus eugenii). Genetics 162:321–330
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754
Zickler D, Kleckner N (2015) Recombination, pairing, and synapsis of homologs during meiosis. Cold Spring Harb Perspect Biol 7:a016626
Acknowledgements
AR-H is supported by the Ministry of Economy, Industry and Competitiveness (CGL2017-83802-P) and the Spanish Ministry of Science and Innovation (PID2020-112557GGB-I00). PDW is supported by the Australian Research Council (DP170101147, DP180100931, DP210103512 and DP220101429). The authors are grateful to C. Vara for providing the first draft for Fig. 1A, C.
Author information
Authors and Affiliations
Contributions
AR-H and PDW conceived, designed and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Associate editor: Barbara Mable.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ruiz-Herrera, A., Waters, P.D. Fragile, unfaithful and persistent Ys—on how meiosis can shape sex chromosome evolution. Heredity 129, 22–30 (2022). https://doi.org/10.1038/s41437-022-00532-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41437-022-00532-2
This article is cited by
-
Divergence and conservation of the meiotic recombination machinery
Nature Reviews Genetics (2023)
-
Mendel’s laws of heredity on his 200th birthday: What have we learned by considering exceptions?
Heredity (2022)