Abstract
The present taxonomic status of Schenkia elegans, endemism recently described in the Iberian Peninsula, and its relationship with the sympatric and the nearest morphological species Schenkia spicata and Exaculum pusillum is revaluated. Different kinds of evidence based on plant morphology, ploidy estimation by flow cytometry, karyotype characterisation, and phylogenetic data have been analysed. Two maternally inherited plastid DNA regions (trnL intron and trnL-F spacer) and biparentally inherited nuclear ribosomal DNA sequence region (nrDNA ITS) have been used. Comparative multivariate analyses show an intermediate morphology of the S. elegans plants between the other two species studied. Flow cytometry and karyotype analyses in S. elegans point to an allopolyploid origin, with the latter constituted by a mixture of those of the diploids S. spicata and E. pusillum. Phylogenetic analyses based on plastid and nuclear DNA regions cluster S. elegans in two different clades, those of S. spicata and E. pusillum, suggesting a possible hybrid origin of S. elegans between both species, acting as maternal or paternal progenitors. In consequence, taking in consideration the taxonomic relationships among genera (Exaculum, Schenkia and the closely related genus Zeltnera found in America), a monotypic genus Valdesiana gen. nov. is proposed to accommodate the allopolyploid species, combined as V. elegans, for which immediate conservation measures must be evaluated.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hybridization in plants is a highly important evolutionary process that favours adaptation to new or unstable habitats, as well as speciation, through the rupture or reinforcement of the reproductive barriers (Grant 1981; Rieseberg 1997; Whitney et al. 2010). Thus, interspecific hybridization could probably represent a source of adaptive genetic variation rather than mutation (Abbott et al. 2013). Hybridization can act in opposition to divergence, introducing an adaptive variation into a population, driving the development of stronger reproductive barriers, or generate new lineages and often creating reticulation patterns (Goulet et al. 2017).
Genome sequence analyses demonstrated that plant hybridization is an ancient evolutionary process (Alix et al. 2017), in which life history, pollination syndrome, breeding system, environment disturbance and genetic predisposition can be important drivers (Ellstrand et al. 1996). Whitney et al. (2010) showed that families and genera differed in hybridization propensity, being particularly common in rich and rapidly diversifying groups of organisms (Seehausen 2013), perhaps because of many young species in geographical proximity.
Hybridization occurs more frequently at the interspecific level. Nevertheless, many cases of intergeneric natural hybridization have been detected in several families of angiosperms (Webb and Druce 1984; Crawford et al. 1993; Weiblen and Brehm 1996; Saito et al. 2006; Soejima et al. 2008; Wu et al. 2010; Cetzal-Ix et al. 2012; Medan et al. 2012; Calvo et al. 2013; Desjardins et al. 2015; Smissen et al. 2015; Anghelescu et al. 2021; Onofre de Araujo et al. 2021). Whitney et al. (2010) found 3.5% of intergeneric hybrids in 3437 hybrids from 13 families, with a ratio higher in those groups with a high number of species.
Hybridization can occur between species sharing or not the same ploidy level, and in the latter, it is frequently accompanied by the subsequent full genome doubling, the hybridization between species or genetically differentiated populations of a species followed by genome duplication (allopolyploidy) could have important genetic and evolutionary consequences (Mable 2004; Mallet 2007; Soltis et al. 2009, 2016; Renny-Byfield & Wendel 2014; Sehrish et al. 2015; Fowler and Levin 2016). In any case, rapid chromosomal evolution and the availability of a suitable habitat for hybrids could favour hybrid speciation (Rieseberg 1997).
Genome duplications can produce a rapid process of reproductive isolation of the progenitors without the need for allopatry because they create a strong, although often incomplete, postzygotic reproductive barrier, and at the same time allowing the restoration of fertility more easily (Grant 1981; Otto and Whitton 2000; Coyne and Orr 2004; Rieseberg and Willis 2007). Likewise, polyploidy plays a predominant role in bursts of adaptive speciation and represents an important source of evolutionary novelty (Fowler and Levin 2016; Alix et al. 2017), by reproductive isolation, morphological differentiation, and deep effects on subsequent lineage evolution (Wood et al. 2009; Te Beest et al. 2012). These changes could be considered as revolutionary (Soltis et al. 2016), if both genetic and epigenetic changes are included. As a result, adaptation to new habitats and geographic areas is expected, with subsequent interactions with herbivorous animals and pollinators, thereby contributing to the emergence of new species (Stebbins 1950, 1985, 2014; Ramsey and Schemske 1998; Soltis et al. 2004; Thompson et al. 2004; Adams and Wendel 2005).
The frequency of genome duplication events throughout the history of seed plants and angiosperms attests to the important role that polyploidy has played in the evolution and diversification of plants in almost all vascular plant lineages (Soltis et al. 2009; Jiao et al. 2011; Wendel 2015; Barker et al. 2016; Alix et al. 2017; Sharbrough et al. 2017), especially in the case of allopolyploidy (Levin 1983; Coyne and Orr 2004; Mallet 2007). This could be explained because allopolyploids are more easily recognizable due to their phenotypic and molecular intermediacy or alternatively by the observation of transgressive phenotypes (Laport and Ng 2017).
The genesis and evolution of allopolyploids could be intricate. Many studies in species of Spermatophyta have evidenced that allopolyploidization events can be dynamic, repeated and that allopolyploid species may have multiple independent origins, as it has been proposed for several polyploids (Govindarajulu et al. 2011; Sigel et al. 2014; Neubig et al. 2015; Vallejo-Marín et al. 2015; Welles and Ellstrand 2016). At present, allopolyploidy represents an important main mechanism of diversification, with several new allopolyploids having originated just within the past century (Soltis et al. 2009).
The complex geological and climatic history of the Mediterranean region is key in favouring speciation processes (Thompson 2005), allowing the alternation of periods of isolation and contact between nearby species, thereby causing gene flow and genetic drift that increase the rate of diversification. In many cases, speciation has been accompanied by hybridization and polyploidy events (Vilatersana et al. 2000), which have been important sources for diversification in some Mediterranean lineages, especially in the Iberian, Italian and Balkan Mediterranean peninsulas (Sáinz Ollero and Moreno Saiz 2002; Vargas et al. 2009; Moreno Saiz 2011; Spaniel et al. 2011; Escudero et al. 2018). In the Iberian Peninsula, 13% of all taxa are considered as hybrid plants, and a 48.8% overall frequency of polyploidy have been reported in several genera (Marques et al. 2018).
In Mediterranean Gentianaceae polyploidy is a source of genetic diversity in genus Centaurium, with 15 polyploid taxa of the 27–30 recognized recently (Jiménez-Lobato et al. 2019), half of them or more suggested to have been originated by allopolyploidy (Ubsdell 1976; Mansion et al. 2005; Guggisberg et al. 2006).
Whitney et al. (2010) indicated for Gentianaceae a low propensity for hybridization, ranked in tenth place out of the 11 defined categories, although Löve (1953) previously indicated that the generic diversification in this family has been based in a high degree on allopolyploidy, as the high variation in the basic number of chromosomes between the different groups shows.
A molecular systematic study of the family Gentianaceae, based on the combination of the phylogenetic approach (trnL intron and matK sequence data), with support of morphological and cytogenetic data, concluded that one of the six monophyletic tribes recognized, Tribe Chironieae, is an important assemblage of 23 genera (Mansion and Struwe 2004). One of them, Centaurium Hill, resulted polyphyletic under its old or classic circumscription (Mansion and Struwe 2004), and consequently, genera Gyrandra Griseb. and Schenkia Griseb. were segregated, and a new genus, Zeltnera Mansion, was recognized (Mansion 2004).
The cytogenetic and phylogenetic analyses from nrDNA and cpDNA sequences (Mansion and Struwe 2004; Mansion and Zeltner 2004) indicate that Zeltnera (with 25 species restricted to the New World) constitutes a well-supported monophyletic assemblage, closely related to Schenkia (with five Mediterranean Basin and Australian species) and the monospecific Mediterranean Basin Exaculum Caruel. The three genera share the morphology of the style with the two stigma lobes converging to form a subcapitate stigma and differ from the clear bifid style found in Centaurium (Mansion 2004).
Schenkia and Zeltnera differ in the type of the inflorescence, spiciform cyme in the first, but corymbiform or paniculate cyme in the second (Mansion 2004), whereas the anthers untwisted in Exaculum were the classic character used by many authors in Euromediterranean Floras for distinguishing it from those genera (Díaz Lifante and Valdés 2014).
Diploid level is present in Exaculum and Schenkia, although in the latter it is present only in one of the five species recognized, S. spicata (L.) G.Mans. This species shows a larger distribution area, from Mediterranean Region to West Asia (Mansion 2004), than Exaculum, which is found only in the West Mediterranean area.
However, every species of Zeltnera cytogenetically studied to date is polyploid, many of them showing a high incidence of dysploidy (Mansion and Zeltner 2004). More recently, a new species of Schenkia, S. elegans (Samp.) Z.Díaz, originally revealed as a race of S. spicata, was described (Díaz Lifante 2012), based on plants with both dichasial and monochasial cymoids, and flowers shortly pedicelled with subcapitate stigma. At present, this species has a very scattered area, limited to western Iberian Peninsula.
The current placement of S. elegans in the genus Schenkia is not entirely satisfactory because some morphological features resemble Exaculum (Díaz Lifante 2012), as the flowers pedicellate, not appressed against stem, calyx lobes unkeeled and erect, and usually 4-lobed corolla. Furthermore, the three Iberian species of these two genera, S. spicata, S. elegans and E. pusillum (Lam.) Caruel often cohabit in the same geographical area. In consequence, a hybrid origin for S. elegans could be hypothesised from these two species belonging to different genera. In this paper, morphological, cytogenetical and molecular analyses have been carried out in order to prove the possible hybrid origin for S. elegans, and its taxonomic consequences.
Materials and methods
Morphological analyses of plants
Detailed morphological multivariate analyses were conducted in wild plants of S. elegans, S. spicata and E. pusillum collected in 17 populations from Iberian Peninsula (Table 1, Fig. 1), most of them belonging to Southwestern region, where these taxa often coexist, and North Morocco. Measurements were obtained from five to ten pressed plants per population, except for four populations (Milfontes, Malalien, Abéjar and Águeda) of E. pusillum in which only one plant was measured. Vouchers are kept in herbaria (Table 1). Analyses at population level allowed an adequate choice of diagnostic characters, considering the variation within individuals and populations.
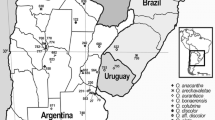
Geographic localization of the populations analysed of Exaculum pusillum, Schenkia elegans and S. spicata. The colour of the dots denotes the species found in the population, in accordance with the legend. The locations have been numbered as indicated in Table 1. See abbreviations for the populations in Table 1.
For examining relationships among the morphological characters and between individual plants, two multivariate ordination analyses were applied to a total of 34 characters (Table 2). A Principal Component Analysis (PCA) was used for 25 continuous characters. For analysing the adequacy of the set of 25 continuous characters to the PCA analyses, both the value for the Kaiser–Meyer–Olkin's parameter, and the significance level for the Bartlett's sphericity test were taken into account. An Optimal Scaling Analysis (OSA) was carried out for nine categorical characters. Analyses of variance (ANOVA) were made among the three taxa by comparing the mean values for each population of the continuous characters, for which previously normality and homoscedasticity were tested with Shapiro–Wilk's and Levene tests (p < 0.05). The multiple comparison post hoc tests of Bonferroni or T2-Tamhane (p less than 0.05), according to the homogeneity of variance, were used for comparing pairs of taxa. The statistic software IBM SPSS Statistics 26.0 (IBM Corp. 1989, 2019) was used for all the analyses.
Furthermore, a morphological comparative analysis was made between plants of S. elegans and the type specimens of several species of Schenkia and Zeltnera, kept in the herbaria BM, BR, E, GH, HAL, JE, K, NY, P and PO [acronyms according to Thiers (2016)].
Cytogenetic analyses
Karyotype analyses
Chromosome counts were carried out in four populations of S. elegans, four populations of S. spicata and four populations of E. pusillum (see Table 1), for which genome size estimates were also obtained. Seeds were collected in several plants from natural populations and sown from January to March at room temperature in Petri dishes containing agar liquid as growth medium (0.125%). Previously, the seeds were surfaced sterilized by soaking them first for 1 min in a solution 10:1 distilled water/bleach (5.5% Sodium hypochlorite), and then, for 1 min in a solution of sterile distilled water. Root (sometimes stem) apical meristems of several seedlings from 5–10 parent plants were used for the somatic chromosome counts. Karyological method follows Díaz Lifante et al. (2009) except for the break of the mitotic metaphase, which was carried out by immersion of seedlings in distilled water at 0–1 °C for 12 h. The best well-spread metaphase plates were photographed with a Leika camera applied to a Zeiss Axioscope microscope at a magnification ranged to 1800.
Conventional karyograms, with chromosome pairs arranged from largest to smallest, first by size and then by asymmetry, did not provide substantial diagnostic information. This is probably due to the very small size and the little differentiation among chromosomes, both in size and in asymmetry, which are gradually reduced. By this reason, the chromosomes for each taxon and population have been distributed in groups according to the length and asymmetry, from highest to shortest values, which facilitated the comparison of karyotypes.
Measurements of the chromosomes were made in the best plates obtained in the different populations studied: seven cells of S. spicata from two populations, eight cells of S. elegans from eight populations and nine cells of E. pusillum from four populations. The ImageJ 1.50i software (Wayne Rasband, National Institute of Health, USA) was used for it. The nomenclature of Stebbins (1938) and Levan et al. (1965) for the apparent size and morphology of chromosomes, respectively, was followed. The karyotype asymmetry was evaluated for each karyogram following Stebbins (1971), and by the indexes A1 (intrachromosomal asymmetry, i.e. based on the ratio of the two arms) and A2 (interchromosomal asymmetry, i.e. based on the chromosome size), as defined by Romero Zarco (1986).
Descriptive statistics (mean and standard deviation of the mean) were calculated for the genome size, the chromosome length (addition of the two chromosome arms), the ratio between the large to short arm of chromosome, and the A1 and A2 indexes of the karyotype asymmetry in each taxon. Univariate comparisons among the three taxa of the mean chromosome length, the total absolute length of the complement, the total length of complement referred to the haploid level and the A1 and A2 asymmetry coefficients were made.
Flow cytometry analysis
DNA content and ploidy level estimates were obtained using propidium iodide flow cytometric (FCM) analysis of nuclei isolated from silica-dried leaf tissues. A total of 119 samples, 33 for E. pusillum from seven populations, 35 for S. elegans from six populations, and 51 for S. spicata from 11 populations were analysed, including in average five plants per population (Table 1). Fresh young leaves were collected, stored in hermetic plastic bags and kept in c. 4 °C during some hours. Once at the laboratory, the samples were dried with silica-gel. Measurements of the DNA content were made in samples of approximately one month old. Nuclei were isolated in a Petri dish containing 1 mL of WPB buffer (0.2 M Tris⋅HCl, 4 mM MgCl2⋅6H2O, 1% Triton X-100, 2 mM EDTA Na2⋅2H2O, 86 mM NaCl, 10 mM metabisulfite, 1% PVP-10; pH adjusted to 7.5 and stored at 4 °C; Loureiro et al. 2007) following the method described by Galbraith et al. (1983). For that, approximately 0.5 cm2 of leaf tissue of our sample was chopped simultaneously with an equal amount of fresh leaf tissue of an internal standard using a razor sharp blade. The reference standard selected was Bellis perennis with a genome size of 2C = 3.57 pg calibrated with Solanum lycopersicum (2C = 1.96 pg, Doležel et al. 1992). Nuclear suspensions were filtered through a 50 µm nylon mesh and stained with 50 µg mL-1 of propidium iodide (PI, Fluka, Buchs, Switzerland). Then, 50 µg mL-1 of RNAse (Fluka, Buchs, Switzerland) was added to avoid staining of double stranded RNA. After 5-min incubation at room temperature, the relative fluorescence intensities of at least 1300 particles per G1 peak were analysed in a Partec CyFlow Space flow cytometer (532 nm green solid-state laser, operating at 30 mW; Partec GmbH, Münster, Germany). The results were obtained using FloMax software (Partec GmbH, Görlitz, Germany) as follows: fluorescence pulse integral in linear scale (FL) histogram; FL vs. time scatter-plot (to assess fluorescence stability); FL vs. fluorescence pulse height scatter-plot (to remove duplets); and FL vs. SS in log scale scatter-plot (to evaluate effects of secondary metabolites; Loureiro et al. 2021). Polygon regions were defined in the FL vs. SS scatter-plot and further applied to the other plots to remove debris (Loureiro et al. 2021). Histograms were evaluated, retaining only samples with a coefficient of variation (CV) below 8% (silica-dried material often generated histograms of lower quality Čertner et al. 2021; Suda and Trávníček 2006); samples with higher CV values were discarded, and a new sample was prepared. The mean (± SD) CV values of the samples retained in this study were of 5.5 ± 1.1 (with minimum of 3.4 and maximum of 8.0).
A proxy of the holoploid genome size (2C) was calculated for each sample using the following equation: Sample 2C nuclear DNA content (pg) = (Sample G1 peak mean / Reference Standard G1 peak mean) * genome size of the Reference Standard. The nomenclature used for the DNA content and C-value followed Greilhuber et al. (2005), using holoploid genome size for the whole chromosome complement (2C) and monoploid genome size for the DNA content of the monoploid genome (1Cx). The monoploid genome size (1Cx) was calculated by dividing the holoploid genome size (2C) by the inferred ploidy level of each taxon in mass values (pg). Based on the genome size values obtained for the population characterized karyologically, diploids were identified to have genome sizes between 1.00 and 1.30 pg and tetraploids between 2.00 and 2.32 pg. The ploidy level was then assigned to all the populations analysed.
Molecular analysis
DNA extraction and molecular DNA regions
Dry leaf tissues from eight individual (Table 1) were used for genomic DNA extractions using Invisorb ® Spin Plant Mini Kit. Accessions from 27 species representing the main lineages of Tribe Chironieae (Gentianaceae) were downloaded from GenBank (Clark et al. 2016); see Supporting Information, Table S1). Specifically, accessions from the nuclear ITS1 and ITS2 regions and plastid trnL intron and trnL-F spacer were downloaded to be used in phylogenetic analyses. These accession sequences were first published by Struwe et al. (2002).
Sequencing
The nuclear ITS (internal transcribed spacer) DNA regions and the plastid DNA regions trnL intron and trnL-F spacer were amplified following protocols in Jiménez-Lobato et al. (2019). The PCR products were sequenced using Macrogene Europe Laboratory services.
Phylogenetic analysis
The sequences were aligned together with other sequences previously downloaded from GenBank using the software MUSCLE (Edgar 2004). Three ITS sequences of S. elegans showed a large number of additivities (ELE1-Alm, ELE2-Alm and ELE1-Puebla). The software jModelTest2 (Darriba et al. 2015) was used to infer the best possible model of nucleotide substitution (GTR + I + G was inferred as the best model in all analyses). The two matrices (cpDNA and ITS) were analysed using the phylogenetic software MrBayes (Ronquist and Huelsenbeck 2003; Ronquist et al. 2012). The analyses were run for 5 million of iterations, and the first 25% of the iterations were discarded as burn-in. The analyses were run twice using for MCMC chains to ensure we reach stationarity and convergence. The post-burn-in trees were used to build a consensus tree. The posterior probability (PP) was used as a measurement of clade support (Alfaro et al. 2003). Finally, an analysis combining the nuclear and plastid matrices was performed with the same parameters but for 10 million iterations.
Results
Plant morphology
Several morphological features in plant, leaves, flowers and fruits have been useful for distinguishing among S. spicata, S. elegans and E. pusillum after the multivariant analyses carried out, many of them shown in Fig. 2.
Distinctive features among Exaculum pusillum, Schenkia elegans and S. spicata (from left to right): a branching pattern of the stem; b flowers, c; pedicel and calyx; d capsule; e half twisted anthers of E. pusillum; f fruiting flower and top vision of flower of S. elegans; g flower of S. elegans showing proximity between anthers and style. Scale bars: b and c, 2.5 mm; d, 2 mm; e, 0.4 mm
The indicators of the adequacy of the PCA are optimal for its application to the entire set of 25 continuous characters (values of 0.927 for the Kaiser–Meyer–Olkin's parameter and high significance for Bartlett’s sphericity test, χ2 = 3835.31, p < 0.0001). Communality was equal to or higher than 0.7 in 22 characters. Five components reach eigenvalues greater than unity (S2), and the first three of them explain 69.40% of the variance. The first component achieves a high eigenvalue, retaining 55.03% of the variance, equivalent to almost 14 characters. In this component, 17 characters have eigenvector larger than 0.7, 12 of them with eigenvalues larger than 0.8. The length of the flower pedicel is the only character that correlates negatively with the rest. The other components retain little variance (< 8%). In the second component, the width of the stigma shows the higher eigenvalue, followed by the width of the lobes, and the length of the seeds and the first cyma internode. The width of the sepals shows eigenvectors values larger than 0.7 in the fourth component.
The two first principal components allow distinguishing the plants belonging to each taxon, placing them in three clearly discontinuous groups (Fig. 3a). No morphological distinction is found for any taxon amongst the plants belonging to populations of different geographical regions. The first component represents many characters relative to the magnitude of different parts of the plant (plant height, leaf length and width, and length of sepals, corolla tube and lobes, stamens, anthers, ovary and pistil), many of them with high weight (up to 0.8) (Figs. 2a–c, e and 3b). This component allows differentiating S. spicata from S. elegans and E. pusillum, which shows the highest values and the greatest variance among the three taxa. Conversely, the lowest values for these characters are found in E. pusillum, whereas S. elegans shows an intermediate position, close to the origin of abscissa axis. The second component, based in fewer characters (seed size, width of corolla lobes and bracts), clearly separates S. elegans from E. pusillum, and rather well from S. spicata (Figs. 2b–c and 3b).
Results of the Principal Component Analysis based on the first two principal components (55.03% and 7.97% of variance): a ordination for the plants of the 12 populations of Exaculum pusillum (1), Schenkia elegans (2) and S. spicata (3); b ordination for the 25 continuous characters. See abbreviations for localities and characters in Tables 1 and 2, respectively
The OSA carried out with the nine categorical characters found values of 0.954 and 0.821 for the Cronbach's Alpha coefficients in the first and second dimensions, with, respectively, the 72.94% and 42.12% of the variance (S3). Three distant groups are differentiated in the ordination of the plants by the two dimensions originated, each one corresponding to a different taxon (Fig. 4a). Dimension 1 separates the populations of S. spicata, whereas dimension 2 distinguishes the populations of S. elegans from that of E. pusillum. The relationships of dependence and independence among the characters analysed show that the roughness and symmetry of the calyx, the arrangement of flowers and the shape of the corolla lobes were very discriminant in the first dimension (Figs. 2b–c and 4b). The apex of the lower leaves and the colour and number of the corolla lobes show medium values in the dimension 2 and somewhat lower values in the first dimension. However, two characters, presence of keel in sepals and number of helical turns of the anther, reached similar and the highest values in the two dimensions (Fig. 2c, e–g).
Results of the Optimal Scaling Analysis (OSA) based on the first two dimensions (72.94% and 42.12% of variance): a ordination of the 12 populations; b ordination of the states of the nine categorical characters; exclusive states for the characters in each taxa are grouped (ELE Schenkia elegans, EXA E. pusillum, SPI S. spicata); shared states are grey-shaded. See abbreviations of localities and characters in Tables 1 and 2, respectively
A great inter-population homogeneity is found in S. elegans, whereas E. pusillum shows the highest dispersion of population data, with Atalaya (ATA) as the most variable population for the features analysed (4–5 dark or light pink corolla lobes) (Fig. 4a). The ordination of the states for the characters shows those exclusive for E. pusillum, as lower leaves acute, usually four corolla lobes (rarely five), which are white-yellowish or pale rose, and anthers non or with a helicoidal turn (Fig. 4b). Five states are exclusive for S. spicata: flowers appressed against to the cyma axis, unequal, twist and smooth sepals (Fig. 2c), corolla lobes acute or somewhat denticulate and anthers with 3–4 helicoidal turns. Schenkia elegans shares four states with E. pusillum (shaded in clear grey in Fig. 4b): obtuse lower leaves, erect flowers, corolla lobes subobtuse and relatively mucronulate, and equal, straight and more or less scabrid sepals (Fig. 2c, f–g). Three states are similar between S. elegans and S. spicata: usually 5 corolla lobes, which are subobtuse, and dark pink to fuchsia (shaded in dark grey in Fig. 4b). The only state exclusive for S. elegans was 1–2 helicoidal turns of the anthers, which really is intermediate between that of the other two taxa. Besides in S. elegans, the upper middle of the calyx lobes detaches from the corolla tube, whereas in S. spicata the calyx lobes applied totally, and in E. pusillum the character is variable, although this feature has not been included in the multivariate analysis.
The ANOVA for the morphological characters found significant differences among the population means at p < 0.0001 (2, 18 d.f.) for 23 of the 25 continuous characters of the three taxa (S4). However, no significant differences at p < 0.05 were found in the length of the first internode of the cyma and the width of sepals. Post hoc Bonferroni tests did not show significant differences at p < 0.05 between S elegans and E. pusillum in three characters (degree of branching, length of plant and length of cyma), and between S. spicata and S elegans in five characters (width of bract and corolla lobe, length of flower pedicel, and finally the fruit and seed).
However, the differences between the three taxa were significant at p < 0.01 in 10 characters (representing 40% of them), both for characters smaller than 2 mm and for characters greater than 2 mm (S5a and S5b, respectively).
Cytogenetic analyses
Chromosome number
The somatic chromosome number 2n = 20 is found in every seedling studied of E. pusillum belonging to 17 parent plants of four populations: three individuals from ALM0, five individuals from ATA, four individual from EMB, and five individuals from DEH) (Fig. 5a-b). In S. spicata, 2n = 22 is counted in the seedlings of 13 parent plants from two populations (eight individuals from ALM0, and five individuals from LIL) (Fig. 5c-d). In S. elegans, a total of 33 parent plants from four populations (four of which from ALM0, twelve coming from ATA, six belonging to DEH and eleven from UPO) are studied, and 2n = 42 is found in every seedling (Fig. 5e-g).
Chromosome size
The mean chromosome length reaches the highest value, 1.862 μm, in E. pusillum, followed by S. spicata with 1.709 μm, and S. elegans with 1.678 μm, but differences are not significant at p < 0.05 (Table 3). The total absolute length of the complement was 35.25 μm in S. elegans, almost twice as that of the other two taxa as can be expected from its polyploid condition (Table 3), although when this length is referred to the haploid level, differences are not found at 0.05 level (F = 0.662, p = 0.526; df = 2).
Karyotype morphology
According to the length and asymmetry of the chromosomes, three groups of them can be distinguished in the karyograms of the three species (Fig. 6). The first group is formed by large to medium sized metacentric (M-m) chromosomes (five pairs in E. pusillum, six pairs in S. spicata and 11 pairs in S. elegans). The second group includes three large to small sized submetacentric (sm) chromosome pairs, both in E. pusillum and S. spicata, and six of them in S. elegans. The third group is formed by two small sized pairs of metacentric (M-m) chromosomes, both in E. pusillum and S. spicata, yet four pairs in S. elegans. In several cells of S. spicata and S. elegans, one or two of the smallest chromosomes of the third group quite often show the two arms widely separated, probably due to the existence of a NOR region near to the centromeric region (see Fig. 5c–d, g). In consequence, the karyograms obtained in S. elegans seem to contain all the chromosome pairs present in those of S. spicata and E. pusillum.
Karyograms for several populations with chromosomes arranged by asymmetry and size in three groups in which they are arranged from largest to smallest: a Exaculum pusillum, 2n = 20 (1, Almendro; 2, Atalaya; 3, Dehesa Abajo; 4, Embalse Grande), b Schenkia spicata, 2n = 22, (1, Almendro; 2, Lillo), c S. elegans, 2n = 42 (1, Almendro; 2, Atalaya; 3, Montequinto). Scale bars: 5 µm (a–c)
Karyotype asymmetry
Following classification of Stebbins (1971), the mean value for the length of the largest and the shortest chromosome pairs in the karyotype of the cells analysed are, respectively, 2.23 and 1.38 μm for E. pusillum, 2.19 and 1.35 μm for S. spicata, and 2.06 and 1.20 μm for S. elegans (S6). Thus, the range for the ratio between the length of the largest to the shortest chromosome is 0.478–0.665 in E. pusillum, 0.489–0.699 in S. spicata and 0.487–0.714 in S. elegans. These values meet with the subtype A2 of chromosomal asymmetry defined by Stebbins (1971), which point out 10–20% of the chromosomes with r ≥ 2 in E. pusillum, 10% in S. spicata and 4.8–9.5% in S. elegans. Furthermore, regarding to the more precise asymmetry indexes A1 and A2 (Romero Zarco 1986), the ANOVA test finds only significant differences for A2 (F = 4.504, p = 0.024; df = 2), although the post hoc Bonferroni test points out significant differences (p = 0.038) only between E. pusillum and S. elegans, with higher values in the latter (Table 3), whereas S. spicata does not distinguish from them.
Flow cytometry analysis. The mean values for the holoploid genome size found are 1.084 ± 0.064 pg in E. pusillum, 1.116 ± 0.039 pg in S. spicata and 2.098 ± 0.053 pg in S. elegans (Table 4). These plants are, thus, classified as having “very small” (2C ≤ 1.4 pg) and “small” (2C ≤ 3.5 pg) genome sizes following Leitch et al. (1998). Significant differences amongst taxa were obtained for the holoploid genome size (F = 325.63, p < 0.0001, df = 2), with S. elegans having significantly higher genome sizes than E. pusillum and S. spicata (p < 0.05). Exaculum pusillum and S. spicata have similar holoploid genome sizes (p > 0.05), and the diploid level is inferred for both taxa. S. elegans present holoploid genome sizes with almost the double of the other two taxa for every population studied (Table 3), and it is inferred as a polyploid, most likely a tetraploid. Assuming these ploidy levels, no significant differences (F = 3.165, p = 0.063, df = 2) are found among taxa in the monoploid genome sizes (0.542 ± 0.032 pg in E. pusillum, 0.556 ± 0.022 pg in S. spicata and 0.524 ± 0.013 pg in S. elegans, Table 3).
Phylogenetic molecular analyses
The new accessions for the plants here studied and obtained in this research are numbered as OQ848573 to OQ848580 for ITS, and OQ851735 to OQ851742 for trnL-F in GenBank. In the ITS tree (S7), the samples SPI-Alm (S. spicata from El Almendro) nests with the sample S. spicata from GenBank (PP = 1) and the sample EXA-Alm (E. pusillum from El Almendro) nests with E. pusillum from GenBank. The sample ELE1-UPO, ELE2-UPO and ELE3-UPO (all of them of S. elegans from the UPO population) groups with EXA-Alm (E. pusillum from El Almendro, PP = 0.93). The samples ELE1-Alm, ELE2-Alm and ELE1-Puebla (S. elegans from El Almendro and Puebla de Guzmán) are sister to the remaining samples of genus Schenkia (PP = 0.99) and grouped with samples from genus Schenkia (PP = 99). These three samples (ELE1-Alm, ELE2-Alm and ELE1-Puebla) are the ones with high number of additivities. These additivities displayed a pattern congruent with a mixture of ITS sequences from S. spicata and E. pusillum.
In the plastid tree (S8), the samples SPI-Alm (S. spicata) and ELE1-Alm, ELE2-Alm and ELE1-Puebla (S. elegans) are nested all together with S. spicata and S. clementii from GenBank (PP = 1). However, the samples ELE1-UPO, ELE2-UPO and ELE3-UPO (S. elegans from UPO) were nested with the sample EXA-Alm (E. pusillum from Almensilla) and E. pusillum from GenBank (PP = 1).
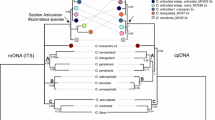
The ITS and plastid combined tree (Fig. 7) displays the simple SPI-Alm clustered in clade A with S. spicata and S. clementii from GenBank (PP = 1). The clade A is sister (PP = 1) to clade B (which includes the samples ELE1-Alm, ELE2-Alm and ELE1-Puebla). The clade C contains the samples EXA-Alm, and E. pusillum from GenBank, which is sister (PP = 1) to clade D (which contains the samples ELE1-UPO, ELE2-UPO and ELE3-UPO, PP = 0.95). The combined tree shows significant support for the relationships of genera Schenkia, Exaculum (including some samples from S. elegans) and Zeltnera. The three genera constitute a monophyletic clade (PP = 1). Genus Zeltnera is sister to the remaining genera (1) and genera Schenkia and Exaculum form a monophyletic clade (PP = 0.95).
Phylogenetic tree of tribe Chironieae, inferred using ITS and plastid sequences. The posterior probabilities (PP) are indicated to the right of the nodes. Clades A, B, C and D are differentiated. Colour squares represent the species: green from Schenkia elegans (ELE), blue for S. spicata (SPI) and S. clementii, and yellow for Exaculum pusillum (EXA). Samples represented with the whole specific name were obtained from the GenBank. For abbreviations of the populations sampled in this research, see Table 1
Discussion
Morphological, cytogenetic, and molecular relationships amongst S. elegans, S. spicata and E. pusillum
Overall, multivariate morphological analyses confirm the recognition of three taxonomic entities clearly distinguishable by numerously continuous and categorical characters, and precise limits can be defined among them. Nevertheless, S. elegans is shown to be intermediate between S. spicata and E. pusillum in many of the morphological characters analysed, which might suggest a hybrid origin between these two species. The continuous traits show a gradient among the three taxa in the size of many structures of the stem, leaves and flowers (Fig. 2a–c), which could be expected for its hybrid nature. However, there are statistically significant clear discontinuities between the three taxa in their ranges of variation for numerous continuous characters, with the highest interpopulation variability found in S. spicata. In relation to the categorical traits, S. elegans shows both parental character states, showing only one intermediate morphological discrete character, the 1–2 helicoidal turns of the anthers, which could be related to the length/width ratio of the anther. S. elegans did not show intermediate but higher values than those present in the other two taxa only in two continuous characters, size of stigma and seeds.
All the above indicates that S. elegans shows the pattern of differentiation that McDade (1990) and Rieseberg and Ellstrand (1993) propose for hybrids, that is, a mosaic of character states present in parents, identical to one or both parents, intermediate between the parental states, more extreme than the parental states, or novel. However, S. elegans shows a low inter-population morphological diversity. As Seehausen (2013) indicated, the recombination of parent genomes after hybridization, causing break down of the genetic correlations, is expected when hybridization is followed by polyploidy. Thus, novel phenotypic characters can represent ‘transgressive segregation’ (Rieseberg et al. 1999) and contribute to the evolution of novel traits and hybrid speciation (Goulet et al 2017). The large stigmas and seeds could favour the self-pollination and seedling establishment, respectively, which furthermore would guarantee the stabilization and perpetuation of the genome of this annual species.
To our knowledge, this is the first estimation of the DNA content for E. pusillum, S. spicata and S elegans. Different DNA amounts between S. elegans and both E. pusillum and S. spicata, which is in accordance with the chromosome numbers found in these species. The chromosome number 2n = 42 found in S. elegans differs from those of S. spicata, with n = 11 and 2n = 22 (results herein and Mesquita Rodrigues 1953; Zeltner 1970, 1991; Carr 1978; Magulaev 1992; Mansion and Zeltner 2004), and E. pusillum, with 2n = 20 (results herein and in Favarger 1960). Interestingly, in other non-Mediterranean species of Schenkia (S. australis, S. clementii and S. sebaeoides), 2n = 44 has been reported (Carr 1978), which indicates a base number of x = 11 for the genus. On the contrary, wide karyological variability exists in the very closely related genus Zeltnera, in which n = 17, 20, 21 and 22 have been indicated, as a result of polyploidy and dysploidy (Broome 1976, 1977, 1978; Turner 1993; Mansion and Zeltner 2004). Moreover, the 2n = 42 of S. elegans agrees with the chromosome number found in some species of Zeltnera belonging to the “Texan group” (Z. maryanna (Turner) Mansion and Z. multicaulis (Robinson) Mansion), and the “Mexican group” (Z. marttinii (Broome) Mansion, Z. nudicaulis (Engelmann) Mansion Z. quitensis and Z. setacea (Bentham) Mansion).
The karyograms reinforce the intermediate karyotype constitution in S. elegans, between that of E. pusillum and S. spicata. Despite S. spicata having two more chromosomes than E. pusillum, no significant differences were found in the DNA amount between E. pusillum and S. spicata, most likely because of the lower size of chromosomes in S. spicata. In S. elegans, the chromosomes are shorter than in S. spicata and E. pusillum, the karyotype shows a higher value for the A2 asymmetry index, and the DNA amount reaches lower values than it can be expected from the correlation between the mean genome size and the ploidy level. After the postpolyploid diploidization process, a “genome downsizing” mechanism, with extensive gene loss, is proposed (Soltis and Soltis 1999; Leitch and Bennett 2004; De Smet et al. 2013; Escudero and Wendel 2020). That mechanism itself could affect the expected correlation between the genome size mean and the ploidy level (Zenil-Ferguson et al. 2016).
Phylogenetic relationships among Schenkia (excluding S. elegans), Exaculum and S. elegans inferred from plastid DNA regions (trnL–intron—trnL-F spacer) revealed two divergent haplotype monophyletic groups for genera Schenkia and Exaculum. The same populations of S. elegans nesting in the same clade, that of S. spicata or that of E. pusillum, do so with both the plastid genome and the nuclear genome, which at the end reinforces their hybrid condition, following Funk (1985). Thus, assuming the maternal inheritance of plastid DNA as in most angiosperms (Smissen et al 2015), both S. spicata and E. pusillum could act as the maternal parent in the hybridization process. In addition, this points out to at least two potential independent evolutionary origins for S. elegans. Until now, S. elegans cohabits with both putative parental species in at least three populations, in which repeated hybridization events between the parent species occur. A similar pattern was found for example by Saito et al. (2006) in the F1 intergeneric hybrid genus Crepidiastrixeris Kitam.
Molecular evidence with nuclear and plastid DNA regions taken together indicates that Exaculum, Schenkia and S. elegans are very closely related and form a monophyletic group which is sister to the North American genus Zeltnera which in time probably has its origins in North America before the Pliocene (Mansion and Struwe 2004; Jiménez-Lobato et al. 2019).
Origin of Schenkia elegans
Every piece of evidence analysed in this study points out to an origin of S. elegans by allopolyploidy, with E. pusillum and S. spicata as the most probable parental species. Although these two diploid species occur in the Mediterranean Basin, the distribution area for E. pusillum (southern and western Europe and northern Africa) is more reduced than that of S. spicata, which spreads up to western Asia, and was recently introduced in North America. The two species are occasionally sympatric in Western Iberian Peninsula (Table 1, Fig. 8a). In three of the populations here analysed, S. elegans grows together with S. spicata and E. pusillum, in other three only with S. spicata, and in one population only with E. pusillum (Table 1). Therefore, S. elegans has probably originated by repeated allopolyploidization at least twice, between the same putative parent species, a situation (polytopic hybrids) well documented and now considered the rule rather than the exception (Soltis and Soltis 1999; Mallet 2007; Leitch and Leitch 2008; Dixon et al. 2009; Soltis et al. 2014; Barker et al. 2016).
The putative diploid parents are morphologically and molecularly well differentiated, which could favour the reproductive stabilization of hybrids after allopolyploidy and the generation of a new species, as Grant (1981) and Stebbins (1985) proposed. Similar cases are found in Gentianaceae, as in some allopolyploid species of Centaurium, such as C. bianoris (Sennen) Sennen, C. malzacianum Maire, and C. mairei Zeltner (Mansion et al. 2005). On the other side, at present there is no evidence for introgression of S. elegans into its putative diploid parents. Furthermore, the great proximity between the anthers and the stigma (Fig. 2 f–g), the large size of this, and the very high fruiting rates (unpublished field observations) suggests that self-pollination may occur in S. elegans, although this must be demonstrated experimentally.
Due to the different diploid genomes of parents, a long-lived perennial growth habit or a self-breeding system is necessary to increase the chance of hybrid individuals producing polyploid offspring (Grant 1981), even more in the case of the allopolyploid species (Stebbins 1950). In consequence, reproductive barriers between polyploids and their diploid progenitors could originate (Rieseberg 1997; Husband and Sabara 2003; Fowler and Levin 2016; Glick et al. 2016; Siopa et al. 2020). Likewise, selfing can decrease the fitness disadvantages in the new polyploids caused by frequency dependent selection (minority cytotype exclusion; Levin 1975) and favours adaptation to new ecological niches or unstable habitats. It represents a reproductive guarantee in cases of low population density and scarcity of pollinators, and furthermore, it would allow safeguarding well-adapted genotypes, as well as their spatial differentiation. A combination of multiple factors related to pre-zygotic and post-zygotic reproductive barriers, and their potentially complex interactions, may also contribute to reproductive isolation (Widmer et al. 2009; Jersáková et al. 2010; Castro et al. 2020), which must be evaluated for S. elegans.
Although somatic doubling and triploid bridge are mechanisms producing polyploidy, the fusion of unreduced gametes from different genomes has been recognized as the most frequent mechanism enabler to allopolyploidy (Fowler and Levin 2016). The rate of production of unreduced gametes increases at higher levels of divergence between hybridizing parents, as does the fertility of polyploid hybrids (Mable 2004; Chapman and Burke 2007; Leitch and Leitch 2008).
The molecular analyses here enclosed show strong relationships among S. spicata, S. clementii, S. elegans and E. pusillum, with the morphologically nearest genus Zeltnera, when ITS and plastid DNA regions are combined. The divergence of Zeltnera from the clade Exaculum-Schenkia is dated by Mansion and Struwe (2004) in an earlier date (higher than 9.2 Mya) than that suggested by Jiménez-Lobato et al. (2019) (late Miocene, c. 7 Mya), during the cooling and aridification of the global climate (Retallack 2004). Following Jiménez-Lobato et al. (2019), Schenkia and Exaculum separated shortly before the Messinian Salinity Crisis, about c.5 Mya. For Mansion and Zeltner (2004), the chromosome number of the species of Zeltnera agrees with a tetraploid constitution, probably originated from unknown diploid ancestors with 2n = 22 and 2n = 20, followed by chromosome rearrangements generating an intense dysploidy series (n = 21 to n = 17). Thus, Zeltnera seems to follow the evolutionary trend proposed by Levin (2020), in which initial polyploidy was followed by a wave of dysploid species formation. Although primary chromosome numbers meet with those present in S. spicata and E. pusillum, the origin of Zeltnera from hybridization between Schenkia and Exaculum does not seem possible, due to the very distant and disjunctive distribution areas of these taxa at present, and the relatively old origin of Zeltnera which predates the divergence of Schenkia and Exaculum (Mansion and Zeltner 2004; Jiménez-Lobato et al. 2019). On the contrary, the species S. elegans seems to have originated in more recent times, showing great morphological similarity between individuals from different populations. Thus, Zeltnera and S. elegans represent examples of the impact of polyploidy on plant diversification by its substantial contribution to the cladogenesis, as indicated by Wood et al. (2009), but in the case of Zeltnera a large later morphological and ecological diversification has been paired by with wide changes in the number of chromosomes.
The plastid sequences found in S. elegans are almost identical to those found in S. spicata or E. pusillum, and the ITS sequences found in S. elegans are almost identical to ITS from E. pusillum or similar to ITS from S. spicata including additive positions congruent with ITS mixing pattern with E. pusillum. These sequences with high number of additivities suggest that convergent ITS evolution is incomplete (which also suggests a very recent origin of this allopolyploid species), but more research is necessary to prove it.
Taxonomic revision
The names of the taxa must be given appropriately to stabilize nomenclature, so that it fulfils its main objective, which is to improve understanding of plant diversification processes and facilitate knowledge for conservation practices of a threatened or endangered singular taxon (Entwisle and Weston 2005; Knapp et al. 2004). Furthermore, the recognition of polyploids as taxonomic entities allows analysing their role in the evolution and in the ecological population dynamic (Laport and Ng 2017), having an impact on a better understanding of biodiversity and its conservation.
Taxonomic classification of polyploid complexes is difficult because their phenotypic differences ranging from subtle to markedly distinct, despite of generally strong reproductive incompatibilities. Unlike autopolyploids, allopolyploids are often well delimited, by their phenotypic and molecular intermediate characters, demonstrating an ecological, geographical and reproductive isolation, by which they deserve taxonomic recognition.
In the present taxonomic treatments, Schenkia and Exaculum constitute two genera well distinguishable morphologically and cytogenetically (Mansion 2004; Mansion & Struwe 2004) One of the five recognized species of Schenkia, S. spicata (= Centaurium spicatum (L.) Fritsch ex Janch.), is the most widely distributed and the only diploid. Much earlier Caruel (1886) described genus Exaculum. Since then, Exaculum has been widely recognized as a monospecific genus in all contemporary Euro-Mediterranean floristic works.
Mansion and Struwe (2004) and Mansion (2004) found that Schenkia and Exaculum diverged several million years ago (see also Jiménez-Lobato et al. 2019), and they differentiated Schenkia from the new genus Zeltnera by the inflorescence. Later, Díaz Lifante (2012) recognised Centaurium spicatum raça elegans Samp. at specific level as Schenkia elegans (Samp.) Z. Díaz, for plants with dichasial and monochasial cymoids, flowers shortly pedicelled and undivided style under the subcapitated stigma. The morphological and cytogenetic analyses showed in the present work point out S. elegans as a well-differentiated taxonomic entity, most likely originated by allopolyploidy from S. spicata and E. pusillum, as molecular data showed, which makes necessary to re-evaluate its taxonomic rank. Furthermore, S. elegans is very probably a fertile entity, as it has already commented above.
Morphological differentiation among S. elegans and the nearest phylogenetic genera
Exaculum differs from Schenkia by the paniculate inflorescence, constituted by dichasial or monochasial cymoids, with flowers largely pedicellate, very short and untwisted after anthesis anthers, rarely somewhat twisted, calyx lobes unkeeled and usually 4-lobed corolla. However, in Schenkia inflorescence is usually spiciform with flowers sessile or subsessile (although in some species as S. clementii is clearly pedicelled), twisted anthers, keeled calyx and 5-lobed corolla. In Zeltnera, genus sister to Exaculum and Schenkia, the anthers are always twisted, but a great morphological variability is found in the other characters: different shape and size of leaves, type of the inflorescence and cymes, length of pedicel, calyx lobes unkeeled or somewhat keeled, corolla 4–5-lobed, and size of the corolla, anthers and style (Mansion 2004). This variability makes difficult to establish generic limits between Zeltnera and these other two genera, although the molecular phylogenetic studies support a clear monophyletic group for Zeltnera (Mansion 2004).
Comparison between S. elegans and the other species of Schenkia
In S. spicata, the monochasial cymoids predominate, with a high number of distant flowers (5–20), sessile or with a very short pedicel, and appressed to the stem. In S. elegans, the stem is branched more often under the main cymoid, dichasial and monochasial cymoids are frequent, there are few flowers per cymoid (2–6) and flowers are distant, clearly pedicellate and divergent. Whereas in S. spicata, the calyx lobes are clearly unequal, bent and appressed to the corolla tube, in S. elegans they are subequal, straight and not fully appressed to the corolla tube. In S. spicata, the corolla lobes are ovate-lanceolate or narrowly elliptic, subobtuse with the apex entire or denticulate and nearly as long as the tube, whereas in S. elegans they are widely elliptic, obtuse, mucronulate and proportionally shorter than the tube. The style is more or less as long as the ovary in S. spicata, but it is clearly shorter in S. elegans.
In S. australis (Brown) Mansion, the flowers are occasionally somewhat pedicellate, the style is shorter than the ovary and the inflorescence is spiciform to racemose (holotype BM 000803703!, isotype K 000750973!). Schenkia clementii (Domin) Mansion has often monochasial cymoids as S. spicata, and longer pedicels (10–20 mm) and flowers (16–18 mm) than in S. elegans, in which, respectively, ranges of 1–2.2(4.5) mm and 7–11 mm are found. Some characters given for Schenkia japonica (Maximovicz) Mansion and Schenkia sebaeoides Grisebach, as wider basal leaves and fleshy leaves, respectively, clearly differentiate them from S. elegans. From a cytogenetic perspective, the genus Schenkia assembles diploid (S. spicata 2n = 22) and polyploid taxa (S. australis, S. clementii, S. sebaeoides) with 2n = 44 (Mansion 2004).
Comparison between S. elegans and Exaculum
Schenkia elegans resembles to E. pusillum in several characters, such as divaricate and often dichotomous branches, cymoids frequently dichasial with a low number of flowers per cymoid, flowers always pedicellate and detached from the stem, calyx lobes somewhat papillose-scabrid subequal and erect, not appressed to the corolla tube, and corolla sometimes 4-merous, with lobes widely elliptic and mucronate. However, in E. pusillum the very short anthers cannot twist after anthesis, and it has smaller flowers and anthers, and narrower leaves than S. elegans.
Comparison between S. elegans and Zeltnera
The high plant morphological variation enclosed in Zeltnera together with the cytogenetic and molecular evidence has allowed to recognize up to 25 species, distributed in three biogeographic groups (Mansion and Zeltner 2004) named as “Californian group”, “Texan group” and “Mexican group”. The analysis of the type specimens for the species of Zeltnera carried out in the present research showed that the plants of S. elegans more closely resemble some species of the “Mexican group”, as Z. quitensis (Kunth) Mansion (holotype:P-670841!; isotype:P-151799!) and Z. stricta (Schiede) Mansion (holotype:HAL-34829!), than those of the “Californian group” or the "Texan group". These two species share with S. elegans the absence of a leaf rosette and the presence of only one well developed stem, broad stem leaves, small-sized flowers (corolla and anthers), a shorter style than the ovary, stigma subcapitate and capsule oblongue to elliptic. However, S. elegans shows the corolla tube longer in relation to the lobes than these two taxa, and the pedicels significantly and persistently much shorter. Furthermore, the chromosome number is variable in both species, with both n = 21 and 22 chromosomes found in each of them (Mansion and Zeltner 2004).
Overall, it is difficult to find diagnostic morphological features for S. elegans that clearly differentiate it from the highly variable genus Zeltnera. Morphologic, cytogenetic and phylogenetic evidence shows that S. elegans is a homogeneous taxon. However, Zeltnera encompasses a myriad of morphological diversity, with a wide combination of traits, as well as a great cytogenetic variability resulting from dysploidy, and a wide biogeographic and ecologic ranges its distribution area. Although S. elegans could be included within the wide morphological diversity of Zeltnera, both taxa have completely different phylogenetic origins, widely separated in time and in different biogeographic contexts.
There are three possible taxonomic solutions here that could reconcile the evolution of the four recognized main lineages: (i) follow a morphological approach and merge Zeltnera, Exaculum, Schenkia and S. elegans in a single genus Schenkia, (ii) merge Exaculum, Schenkia and S. elegans in a single genus Schenkia, or (iii) follow the phylogenetic evidence and describe a new monotypic genus for S. elegans. Following the recommendations of Stuessy (2009) for remodelling genera, to consider an unique genus Schenkia for Zeltnera, Schenkia, Exaculum and S. elegans would be inadvisable from a practical point of view and a theoretical perspective, and it would obligate to make numerous new combinations for the 25 species of Zeltnera, which would increase the list of indexed names in databases and create confusion among the users of classification with conservation purposes. Only one nomenclatural change would occur if Exaculum, Schenkia and S. elegans are including in genus Schenkia. However, in Exaculum a high number of autapomorphies in the flower characters have accumulated, which has increased its morphological differentiation and a high ecological specialization, reinforcing its biological unit. Besides, from a practical point of view for users of classification, there is an old and long tradition of recognizing Exaculum as a highly different and monotypic genus in Mediterranean flora (Díaz Lifante and Valdés 2014). The phylogenetic data (Mansion & Struwe 2004) indicate that this is a very old and distinctive lineage that diverged long time ago from Schenkia and Zeltnera. Moreover, the fact that S. spicata and E. pusillum can hybridise only proves the existence of a common ancestry between both genera, although it does not force to place them in a unique genus. As McKenzie et al. (2004) stated, “A rigid requirement for cross-incompatibility between genera would result in fewer, larger genera that might obscure rather than clarify phylogenetic relationships”.
The three taxa share a wet habitat, but there are certain differences both in the phenology and in the substrate they occupy. Exaculum pusillum is found in community of therophytes in the margin of streams and seasonal ponds, in humid and acid substrates (siliceous sands, slates) which is recognised as Isoeto-Nanojuncetea association. Schenkia spicata has a wider distribution area, also in margin of streams and seasonal ponds, when they have dried. It preferably inhabits basic soils and somewhat salty, as in littoral ponds, but it can be found also on acid substrates. Schenkia elegans occupies the margin of depressions, coastal or inland, as well as the dried channels of ponds and temporary watercourses, on acid (Andévalo, Bajo Guadalquivir), or basic (Beira litoral) substrates. Nevertheless, there is a phenological difference, S. spicata flowers later than S. elegans and E. pusillum. Accordingly, the monotypic genus is the best solution for us and we propose to consider a new genus to accommodate the allopolyploid entity S. elegans. This decision is supported by the existence in S. elegans of an intermediate, but not overlapping, range for many quantitative continuous features which differentiates it clearly from Exaculum and other Schenkia species, two of them exclusive (width of stigma and length of seeds), and one discrete (1–2 helicoidal turns of the anthers).
As a result of this research, and in order to achieve stability for the classification, here a new genus, Valdesiana, is described for the well-established Iberian endemism S. elegans, in the important diversification centre of Mediterranean Region, which could represent the origin of an emerging evolutionary lineage.
Taxonomic treatment
Key to the most phylogenetically related genera
-
1a
Style slightly bifid; capsule subcylindric, oblongoid, or ovoid-oblongoid ……………… Centaurium
-
1b
Style with the two stigma lobes converging to form a subcapitate stigma; capsule ellipsoid to ovoid ……………… 2
-
2a
Anthers not helicoidally coiled, or at most twisted only half a turn ……………… Exaculum
-
2b
Anthers helicoidally coiled after anthesis ……………… 3
-
3a
Inflorescence spiciform, sometimes racemose, often formed by monochasial cymes. Flowers usually sessile to subsessile, rarely pedicellate. Calyx lobes keeled; corolla usually 5-lobed ……………… Schenkia
-
3b
Inflorescence corymbiform, paniculate, racemose or capituliform, formed by both monochasial and dichasial cymes. Flowers subsessile or pedicellate. Calyx lobes unkeeled or slightly keeled; corolla 4–5-lobed ……………… 4
-
4a
Flower pedicels (2) 3-25 (50) mm ……………… Zeltnera
-
4b
Flower pedicels 1-2.2 (4.5) mm ……………… Valdesiana
Valdesiana Z.Díaz & M.Escudero, gen. nov.—TYPE: Valdesiana elegans (Samp.) Z.Díaz & M.Escudero, comb. nov.
Eponymy: The name of the genus is dedicated to Prof. Benito Valdés, eminent and indefatigable Mediterranean botanist who developed important projects of Flora and Taxonomy, and trained a good number of taxonomists at the University of Seville.
Diagnosis: Herb, annual, uniacaule. Basal rosette of leaves absent. Cauline leaves ovate-elliptical or elliptical, acuminate. Flowers (4)-5 merous, not appressed to the stem, arranged in lax and compound dichasial and monochasial cymes. Calyx lobes (4)-5 equal, rarely subequal, somewhat keeled, mucronate. Corolla with (4)-5 lobes, 1/3–1/2 tube length, obtuse, mucronulate, rose-purple, rarely white. Anthers helicoidally coiled in 1–2 turns after dehiscence. Ovary superior, sessile, ovoid-ellipsoid; style marked, 1/2–2/3 of the ovary length, bifurcate very shortly in the upper part; stigma lobes entire, round, convergent in anthesis, subcapitate.
Description: Herbs, annual, uniacaule, with sympodial branching, glabrous. Stem erect, branched in the upper part, sometimes also in the lower part. Basal rossette of leaves absent; middle and upper leaves ovate-elliptical or elliptical, acuminate, with 3–5 nerves. Inflorescence cymose, formed by compound cymes, with dichasial branching in the proximal part and monochasial in the distal part, sometimes with some intercalar dichasial branch, very lax. Flowers pentamerous, rarely tetramerous, almost sessile or shortly pedicelled, separate from the stem. Calyx lobes (4)-5 equal, rarely subequal, not exceeding or exceedingly somewhat the corolla tube, rather keeled, smooth or somewhat scabrid, mucronate, separate and not applied to the corolla tube. Corolla hypocrateriform; lobes 4–5, 1/3–1/2 tube length, entire, obtuse, mucronulate, patent, rose-purple, rarely white. Androecium with 4–5 stamens, erect, moderately converging in anthesis; exceeding or equalling the stigma; anthers slightly sagittate in the base, helicoidally coiled in 1–2 turns after dehiscence. Ovary superior, sessile, ovoid-ellipsoid, and style defined, 1/2–2/3 of the ovary length, slightly bifurcate in the upper part, each branch widened in a stigma lobe; stigma lobes entire, round, convergent in anthesis, subcapitate, with long papilles, yellow, reaching or exceeding somewhat the mouth of the corolla tube. Fruit an ovoid-ellipsoid capsule.
Valdesiana elegans (Samp.) Z.Díaz & M.Escudero, comb. nov. ≡ Centaurium spicatum raça elegans Samp., Man. Fl. Portug.: 383 (1913), basion. ≡ Schenkia elegans (Samp.) Z.Díaz, Flora Iberica 11: 85 (2012).—LECTOTYPE: (designated here): Ílhavo: Ria, 30 Jun 1901, G. Sampaio (PO-6749 G.S. [photo]!). (Fig. 9). Ind. loc.: “Prados maritimos, de Esmoriz a Mira”.
Etymology: The name refers to the slender stems and thinner branches than the type of Centaurium spicatum.
Lectotypification: Sampaio (1913: 383) described Centaurium spicatum raça elegans based on plants from Beira Litoral. He differentiated it from the type subspecies by its slender stems with thin and quite divergent branches, and by its pedicellate and distant flowers, with calyx lobes very similar to each other. Among the materials of Sampaio kept in the Herbarium of Oporto University (PO), the only material identified as Centaurium spicatum raça elegans is the voucher numbered as “6749 G. S.”. It contains four plants measuring 12–17 cm long, in an early flowering state, with 6–7 nodes in the stem, middle leaves 7–12 × 2.5–4.8 mm, cymes monochasial or dichasial with 3–4 nodes, and flowers 9–11 mm, with pedicels 1.2–4 mm, calyx 5.5–7 mm, and corolla with tube 6–7.5 mm and lobes 2.5–3 mm.
Two labels made out of strips of newspaper are included in this voucher (Fig. 9). In the upper label Sampaio wrote “Ílhavo: Ria. 30, 6°. 1901 // Existe tamben em Esmoriz. Tem um porte bem diverso [= very different] da E. spicata. / E' pequena, subglauca”. In the lower label Sampaio wrote “E. spicata / precox, nob. Differe do typo por florescer mais cedo [= early], pelos / ramosi (?) quasi capillares, pelas flores mais distantes, pelos ra- / mos mais aberto-divergentes, pelos” In the lower corner of the right side Sampaio wrote "E. spicata / raç. elegans Samp. // Ílhavo: Ria", using Erythraea, a synonym of the more recent name Centaurium. The standard label of the Instituto de Botánica "Dr. Gonçalo Sampaio" of the University of Porto identifies the specimen, numbered as “6749 G. S.”, as “Centaurium spicatum Frits. rac. elegans Samp.”, collected on “30–VI–1901” in “Ílhavo: Ria” by “Gonçalo Sampaio”. The measurements exhibited by the four plants in the voucher and the Sampaio's handwritten notes (Fig. 9) on the sheet are in agreement with the description given by Sampaio (1913: 383) for this taxon. On page 383 of the Manual da Flora Portugueza, Sampaio indicated a wide area, “Prados maritimos, de Esmoriz a Mira”, where “Ílhavo” is located. In consequence, this voucher contains plants that are type specimens. The first plant on the right, measuring 15 cm high, with a whole stem and at least one mature flower, is chosen as the better plant for representing Centaurium spicatum raça elegans Samp. (Fig. 9).
Chromosome number: 2n = 42 (here published for the first time).
Phenology: Flowering from end-May to late July; mature seeds are available from end-June to August.
Habitat: Temporarily flooded depressions and desiccated streams, coastal or inland lagoons, in brackish or sandy substrates; 0–700 m.
Distribution area: West Iberian Peninsula. Portugal: Beira litoral; Spain: Huelva, Valladolid and Sevilla provinces.
Addition specimens studied: PORTUGAL. Beira Litoral: De Esmoriz a Mira, 30 Jun 1901, G. Sampaio (PO-Samp). SPAIN. Huelva: Puebla de Guzmán, represa cercana al pueblo, 20 Jul 1989, S. Silvestre (SEV-214292). Villanueva de las Cruces, La Tiesa, 6 Jun 2002, C. Santa-Bárbara and Z. Díaz Lifante (SEV-214423). El Almendro, 2 Jul 2010, V. Girón and Z. Díaz Lifante (SEV-249968); ídem, Z. Díaz Lifante and F. García Díaz, 2 Aug 2015 (SEV-270212); ídem, Entre El Almendro y Alosno, proximidades a un curso de agua, 24 Jun 2016, Z. Díaz Lifante and C. García Llamas (SEV 270231). Entre Puebla de Guzmán y El Almendro, 7 Jun 2015, Z. Díaz Lifante (SEV-270205). Sevilla: Coria del Río, “Dehesa de la Atalaya”, 5 Jun 2016, C. Andrés Camacho, Z. Díaz Lifante, J. Díaz Fernández and C. García Llamas (SEV 270230). Puebla del Río, “Dehesa de Abajo”, Z. Díaz Lifante, J. Díaz Fernández and C. García Llamas (SEV 270229). Entre Sevilla y Montequinto, Campus de la Universidad Pablo de Olavide, 1 Jun 2015, Z. Díaz Lifante (SEV-270203). Valladolid: Aldeamayor de San Martín, 26 Jul 1983, M. Ladero, F. Navarro and C. Valle (SALAF-4797).
Conservation status: Not evaluated yet. In previous analyses, Díaz Lifante et al. (2018) indicated a variation in the number of plants per square meter between 0.51 (population of Atalaya, here named as ATA) and 25.9 (population of El Almendro, here named as ALM), with a significant amount of 6.79 in a human disturbed habit at the University Campus of Pablo Olavide. The tests of seed germination here carried out have shown that it is achieved easily in autumn at room temperature, so it seems that there is not a deep or complete dormancy mechanism. But it is urgent to develop a searching program in similar habitats in a more extensive area to complete the knowledge of its distribution, and to monitor populations evaluating its present threat status which would allow for a more accurate categorization of extinction risk (IUCN 2019).
Ecological observations: Plants of V. elegans blooms in early summer, occupying the margins of temporal stream beds, ponds and damp places when dried up and the competition with other herbaceous plants is low. The populations in Huelva and Valladolid provinces colonize soils formed on acid rocks, such as slates and granite, but the populations from Seville province grow in loamy soil.
In the location of Aldeamayor de San Martín, Villanueva de Las Cruces, and Puebla de Guzmán, V. elegans cohabits with S. spicata, but to date only in two locations, El Almendro (Huelva province, Spain) and Dehesa de Abajo (Sevilla province, Spain) E. pusillum is also found (Fig. 8a, b). It is very probable that the three taxa coexist in Beira Litoral province (Portugal), from where herbarium specimens have been studied. In the recent localization in wastelands in the Campus of the University Pablo de Olavide (Seville province), V. elegans occupies depressions retaining moisture for some time, very close to the alluvial terrace of the Guadaira river, near where this flows into the Guadalquivir river. There V. elegans cohabits with S. spicata, but this species flowers one month later.
Data availability statement
All data generated or analysed during this study are included in this published article (and its supplementary information files). The DNA sequences generated during this research will be deposited in the Genbank.
References
Abbott RJ, Albach D, Ansell S, Arntzen JW, Baird SJE, Bierne N, Boughman J, Brelsford SA, Buerkle CA, Buggs R, Butlin RK, Dieckmann U, Eroukhmanoff F, Grill A, Cahan SH, Hermansen JS, Hewitt G, Hudson AG, Jiggins C, Jones J, Keller B, Marczewski T, Mallet J, Martínez-Rodríguez P, Möst M, Mullen S, Nichols R, Nolte AW, Parisod C, Pfennig K, Rice AM, Ritchie MG, Seifert B, Smadja CM, Stelkens R, Szymura JM, Väinölä R, Wolf JBW, Zinner D (2013) Hybridization and speciation. J Evol Biol 26:229–246. https://doi.org/10.1111/j.1420-9101.2012.02599.x
Adams KL, Wendel JF (2005) Polyploidy and genome evolution in plants. Curr Opin Pl Biol 8:135–141. https://doi.org/10.1016/j.pbi.2005.01.001
Alfaro M, Zoller S, Lutzoni F (2003) Bayes or bootstrap? A simulation study comparing the performance of Bayesian Markov chain Monte Carlo sampling and bootstrapping in assessing phylogenetic confidence. Molec Biol Evol 20:255–266. https://doi.org/10.1093/molbev/msg028
Alix K, Gérard PR, Schwarzacher T, Heslop-Harrison (2017) Polyploidy and interspecific hybridization: partners for adaptation, speciation and evolution in plants. Ann Bot (Oxford) 120:183–194. https://doi.org/10.1093/aob/mcx079
Anghelescu NEDG, Kertész H, Constantin N, Simon-Gruița A, Duță Cornescu G, Pojoga MD et al (2021) New intergeneric orchid hybrid found in Romania × Pseudorhiza nieschalkii (Senghas) P.F. Hunt nothosubsp. siculorum H. Kertész & N. Anghelescu, 2020. PLoS ONE 16:e0241733. https://doi.org/10.1371/journal.pone.0241733
Barker MS, Husband BC, Pires JC (2016) Spreading Winge and fl ying high: The evolutionary importance of polyploidy after a century of study. Amer J Bot 103:1139–1145. Available at: http://www.jstor.org/stable/44252765
Broome CR (1976) The Central American species of Centaurium (Gentianaceae). Brittonia 28:413–426. https://doi.org/10.2307/2805605
Broome CR (1978) Chromosome numbers and meiosis in North and Central American species of Centaurium (Gentianaceae). Syst Bot 3:299–312. https://doi.org/10.2307/2418299
Broome CR (1977) Four new species of Centaurium from Mexico. Madroño 24:237–244. http://www.jstor.org/stable/41424116
Calvo J, Alvarez I, Aedo C, Pelser PB (2013) A phylogenetic analysis and new delimitation of Senecio sect. Crociseris (Compositae:Senecioneae), with evidence of intergeneric hybridization. Taxon 62:127–140. https://doi.org/10.1002/tax.621011
Carr GD (1978) Chromosome numbers of Hawaiian flowering plants and the significance of cytology in selected taxa. Amer J Bot 65:236–242. https://doi.org/10.1002/j.1537-2197.1978.tb06061.x
Caruel T (1886) Flora italiana, 6 Corolliflore. Tipographia Dei Successori Le Monnier, Firenze. https://doi.org/10.5962/bhl.title.6341
Castro M, Loureiro J, Husband BC, Castro S (2020) The role of multiple reproductive barriers: strong post-pollination interactions govern cytotype isolation in a tetraploid–octoploid contact zone. Ann Bot (Oxford) 126:991–1003. https://doi.org/10.1093/aob/mcaa084
Čertner M, Lučanová M, Sliwinska E, et al (2021) Plant material selection, collection, preservation, and storage for nuclear DNA content estimation. Cytometry A 101:737–748. https://doi.org/10.1002/cyto.a.24482
Cetzal-Ix W, Balam-Narváez R, Carnevali G (2012) A new nothogenus and nothospecies in the Oncidiinae (Orchidaceae) from Quintana Roo, Mexico. Nordic J Bot 30:40–46. https://doi.org/10.1111/j.1756-1051.2011.01261.x
Chapman M, Burke JM (2007) Genetic Divergence and Hybrid. Evolution 61:1773–1780. https://doi.org/10.1111/j.1558-5646.2007.00134.x
Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2016) GenBank. Nucl Acids Res 44(D1):D67–D72. https://doi.org/10.1093/nar/gkv1276
Coyne JA, Orr HA (2004) Speciation. Sinauer Associates, Sunderland, MA. https://www.joelvelasco.net/teaching/2890/coyneorr04-speciationch1.pdf
Crawford DJ, Brauner S, Cosner MB, Stuessy TF (1993) Use of RAPD markers to document the origin of the intergeneric hybrid xMargyracaena skottsbergii (Rosaceae) on the Juan Fernandez Islands. Amer J Bot 80:89–92. https://doi.org/10.1002/j.1537-2197.1993.tb13771.x
Darriba D, Taboada GL, Doallo R, Posada D (2015) Europe PMC Funders Group jModelTest 2: more models, new heuristics and high- performance computing. Nature Meth 9:772. https://doi.org/10.1007/BF00937587
De Smet R, Adams KL, Vandepoele K, Van Montagu MCE, Maere S, Van de Peer Y (2013) Convergent gene loss following gene and genome duplications creates single-copy families in flowering plants. Proc Natl Acad Sci USA 110:2898–2903. https://doi.org/10.1073/pnas.1300127110
Desjardins DS, Leslie CA, Stace AC, Schwarzacher T, Bailey PJ (2015) Intergeneric hybridisation between Berula erecta and Helosciadium nodiflorum Apiaceae. Taxon 64:784–794. https://doi.org/10.12705/644.9
Díaz Lifante Z, Valdés B (2014) Lectotypification of Gentiana filiformis and Gentiana pusilla (Gentianaceae). Taxon 63:417–422. https://doi.org/10.12705/632.3
Díaz Lifante Z, Andrés Camacho C, Viruel J, Cabrera Caballero A (2009) The allopolyploid origin of Narcissus obsoletus (Alliaceae): identification of parental genomes by karyotype characterization and genomic in situ hybridization. Bot J Linn Soc 159:477–498. https://doi.org/10.1111/j.1095-8339.2009.00951.x
Díaz Lifante Z, García Llamas C, Díaz Fernández J, Andrés Camacho C (2018) Nuevas localizaciones para Schenkia elegans (Samp) Z Díaz (Gentianaceae) en Andalucía Occidental con apuntes sobre su demografía. Acta Bot Malac 43:145–147. https://doi.org/10.24310/abm.v43i0.5314
Díaz Lifante Z (2012) Schenkia Griseb. In: Castroviejo S, Andrés C, Arista M, Fernández-Piedra P, Gallego MJ, Ortiz PL, Romero C, Salgueiro FJ, Silvestre S and Quintanar A (eds) Flora Iberica, 11:81–86. Gentianaceae-Boraginaceae. Real Jardín Botánico, CSIC, Madrid. http://www.floraiberica.es/floraiberica/texto/pdfs/11_130_07_Schenkia.pdf
Dixon JD, Schönswetter P, Suda J, Wielderaminn MM, Schneeweiss GM (2009) Reciprocal Pleistocene origin and postglacial range formation of an allopolyploid and its sympatric ancestors (Androsace affinis group, Primulaceae). Molec Phylogen Evol 50:74–83. https://doi.org/10.1016/j.ympev.2008.10.009
Doležel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol Pl 85:625–631. https://doi.org/10.1111/j.1399-3054.1992.tb04764.x
Edgar RC (2004) MUSCLE:multiple sequence alignment with high accuracy and high throughput. Nucl Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Ellstrand N, Whitkus R, Rieberg L (1996) Distribution of spontaneous plant hybrids. Proc Natl Acad Sci USA 93:5090–5093. https://doi.org/10.1073/pnas.93.10.5090
Entwisle TJ, Weston PH (2005) Majority rules, when systematists disagree. Austral Syst Bot 18:1–6. https://doi.org/10.1071/SB04013
Escudero M, Wendel JF (2020) The grand sweep of chromosomal evolution in angiosperms. New Phytol 228:805–808. https://doi.org/10.1111/nph.16802
Escudero M, Balao F, Martín-Bravo S, Valente L, Valcárcel V (2018) Is the diversification of Mediterranean Basin plant lineages coupled with karyotypic changes? Pl Biol 20 (Suppl 1):166–175. https://doi.org/10.1111/plb.12563
Favarger C (1960) Etude cytologique du Cicendia filiformis et du Microcala pusilla (Gentianacées). Bull Soc Bot France 107:94–98. https://doi.org/10.1080/00378941.1960.10837919
Fowler NL, Levin DA (2016) Critical factors in the establishment of allopolyploids. Amer J Bot 103:1236–1251. https://doi.org/10.3732/ajb.1500407
Funk VA (1985) Cladistics and generic concepts in the Compositae. Taxon 34:72–80. https://doi.org/10.2307/2399220
Galbraith DW, Harkins KR, Maddox JM, Ayres NM, Sharma DP, Firoozabady E (1983) Rapid flow cytometric analysis of the cell cycle in intact plant tissues. Science 220:1049–1051. https://doi.org/10.1126/science.220.4601.1049
Glick L, Sabath N, Ashman TL, Goldberg E, Mayrose I (2016) Polyploidy and sexual system in angiosperms: is there an association? Amer J Bot 103:1223–1235. https://doi.org/10.3732/ajb.1500424
Goulet BE, Roda F, Hopkins R (2017) Hybridization in plants: old ideas, new techniques. Pl Physiol 173:65–78. https://doi.org/10.1104/pp.16.01340
Govindarajulu R, Hughes CE, Alexander PJ, Bailey CD (2011) The complex evolutionary dynamics of ancient and recent polyploidy in Leucaena (Leguminosae; Mimosoideae). Amer J Bot 98:2064–2076. https://doi.org/10.3732/ajb.1100260
Grant V (1981) Plant speciation. Colimbia Univ Press, New York. https://doi.org/10.3732/ajb.1100260
Greilhuber J, Dolezel J, Lysák MA, Bennett MD (2005) The origin, evolution and proposed stabilization of the terms “Genome Size” and “C-value” to describe nuclear DNA contents. Ann Bot (Oxford) 95:255–260. https://doi.org/10.1093/aob/mci019
Guggisberg A, Bretagnolle F, Mansion G (2006) Allopolyploid origin of the Mediterranean endemic, Centaurium bianoris (Gentianaceae), inferred by molecular markers. Syst Bot 31:368–379. https://doi.org/10.1600/036364406777585937
Husband BC, Sabara HA (2003) Reproductive isolation between autotetraploids and their diploid progenitors in fi reweed, Chamerion angustifolium (Onagraceae). New Phytol 161:703–713. https://doi.org/10.1046/j.1469-8137.2004.00998.x
IBM Corp (1989, 2019) IBM SPSS Statistics for Windows, version 26.0. IBM Corp., Armonk. Available at: https://www.ibm.com/support/pages/downloading-ibm-spss-statistics-26
ImageJ 1.50i (Wayne Rasband, National Institute of Health, USA). Available at: http://imagej.nih.gov/ij/
IUCN Standards and Petitions Subcommittee (2019) Guidelines for using the IUCN red list categories and criteria. Version 14. Prepared by the Standards and Petitions Committee. Available at: http://www.iucnredlist.org/documents/RedListGuidelines.pdf
Jersáková J, Castro S, Sonk N, Milchreit K, Schödelbauerová I, Tolasch T, Dötterl S (2010) Absence of pollinator-mediated premating barriers in mixed-ploidy populations of Gymnadenia conopsea s.l. (Orchidaceae). Evol Ecol 24:1199–1218. https://doi.org/10.1007/s10682-010-9356-7
Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L et al (2011) Ancestral polyploidy in seed plants and angiosperms. Nature 473:97–100. https://doi.org/10.1038/nature09916
Jiménez-Lobato V, Escudero M, Díaz Lifante Z, Andrés Camacho C, de Castro A, Mansion G, Zeltner L, Arroyo J (2019) Evolution of reproductive traits and selfing syndrome in the sub-endemic Mediterranean genus Centaurium Hill (Gentianaceae). Bot J Linn Soc 191:216–235. https://doi.org/10.1007/s11033-021-06764-5
Knapp S, Lamas G, Lughadha EN, Novarino G (2004) Stability or stasis in the names of organisms:the evolving codes of nomenclature. Philos Trans Roy Soc London B, Biol Sci 359:611–622. https://doi.org/10.1098/rstb.2003.1445
Laport RG, Ng J (2017) Out of one, many: The biodiversity considerations of polyploidy. Amer J Bot 104:1119–2112. https://doi.org/10.3732/ajb.1700190
Leitch IJ, Bennett MD (2004) Genome downsizing in polyploid plants. Biol J Linn Soc 82:651–663. https://doi.org/10.1111/j.1095-8312.2004.00349.x
Leitch AR, Leitch IJ (2008) Genomicp plasticity and the diversity of polyploid plants. Science 320:481–483. https://doi.org/10.1126/science.1153585
Leitch IJ, Chase MW, Bennett MB (1998) Phylogenetic analysis of DNA C-values provides evidence for a small ancestral genome size in flowering plants. Ann Bot (Oxford) 82:85–94. https://doi.org/10.1006/anbo.1998.0783
Levan AK, Fredga K, Sandberg AA (1965) Nomenclature for centromeric position on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Levin DA (1975) Minority cytotype exclusion in local plant populations. Taxon 24:35–43. https://doi.org/10.2307/1218997
Levin DA (1983) Polyploidy and novelty in flowering plants. Amer Naturalist 122:1–25. https://doi.org/10.1086/284115
Levin DA (2020) Did dysploid waves follow the pulses of whole genome duplications? Pl Syst Evol 306:5. https://doi.org/10.1007/s00606-020-01704-5
Loureiro J, Rodriguez E, Doležel J, Santos C (2007) Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann Bot (Oxford)100:875–888. https://doi.org/10.1093/aob/mcm152
Loureiro J, Kron P, Temsch EM, Koutecký P, Lopes S, Castro M, Castro S (2021) Isolation of plant nuclei for estimation of nuclear DNA content: overview and best practices. Cytometry A 99:318–327. https://doi.org/10.1002/cyto.a.24331
Löve D (1953) Cytotaxonomical remarks on the Gentianaceae. Hereditas 39:225–235. https://doi.org/10.1111/j.1601-5223.1953.tb03415.x
Mable BK (2004) Polyploidy and self-compatibility: is there an association? New Phytol 162: 803–811. https://www.jstor.org/stable/1514575
Magulaev AY (1992) Chromosome numbers in some species of vascular plants of the northern Caucasus flora. Bot Žhur (Moscow & Leningrad) 77(10):88–90. [Ref. is not correct to find]
Mallet J (2007) Hybrid speciation. Nature 446:279–283. https://doi.org/10.1038/nature05706
Mansion G (2004) A new classification of the polyphyletic genus Centaurium Hill (Chironiinae-Gentianaceae): description of the New-World endemic Zeltnera, and reinstatement of Gyrandra Griseb. and Schenkia Griseb. Taxon 53:719–740. https://doi.org/10.2307/4135447
Mansion G, Struwe L (2004) Generic delimitation and phylogenetic relationships within the subtribe Chironiinae (Chironieae:Gentianaceae), with special reference to Centaurium:evidence from nrDNA and cpDNA sequences. Molec Phylogen Evol 32:951–977. https://doi.org/10.1016/j.ympev.2004.03.016
Mansion G, Zeltner L (2004) Phylogenetic relationships within the New World endemic Zeltnera (Gentianaceae-Chironiinae) inferred from molecular and karyological data. Amer J Bot 91:2069–2086. https://doi.org/10.3732/ajb.91.12.2069
Mansion G, Zeltner L, Brettagnole F (2005) Phylogenetic patterns and polyploid evolution within the Mediterranean genus Centaurium (Gentianaceae – Chironieae). Taxon 54:931–950. https://doi.org/10.2307/25065479
Marques I, Loureiro J, Draper D, Castro M, Castro S (2018) How much do we know about the frequency of hybridisation and polyploidy in the Mediterranean region? Pl Biol 20 (Suppl 1):21–37. https://doi.org/10.1111/plb.12639
McDade L (1990) Hybrids and phylogenetic systematics I. Patterns of character expression in hybrids and their implications for cladistic analysis. Evolution 44:1685–1700. https://doi.org/10.1111/j.1558-5646.1990.tb03856.x
McKenzie RJ, Ward JM, Lovis JD, Breitwieser I (2004) Morphological evidence for natural intergeneric hybridization in the New Zealand Gnaphalieae (Compositae): Anaphalioides bellidioides × Ewartia sinclairii. Bot J Linn Soc 145:59–75. https://doi.org/10.1111/j.1095-8339.2003.00282.x
Medan D, Arbteman M, Chaia EE, Premoli AC (2012) Interspecific and intergeneric hybridization in South American Rhamnaceae-Colletieae. Pl Syst Evol 298:1425–1435. https://doi.org/10.1007/s00606-012-0646-0
Mesquita Rodrigues JE (1953) Contribuição para o conhecimento cariológico das halófilas e psamófitas litorais. PhD Thesis, The University of Coimbra, Coimbra
Moreno Saiz JC (2011) La diversidad florística vascular española. Mem Real Soc Esp Hist Nat. 2ª ép., 9:75–107. http://www.rsehn.es/cont/publis/boletines/128.pdf
Neubig KM, Blanchard OJ, Whitten WM, McDaniel SF (2015) Molecular phylogenetics of Kosteletzkya (Malvaceae, Hibisceae) reveals multiple independent and successive polyploid speciation events. Bot J Linn Soc 179:421–435. https://doi.org/10.1111/boj.12330
Onofre de Araujo A, Peixoto M, Neves de Souza C, Custodio Gasparino E, Toledo Faria J, Augusto Lombello R (2021) A natural intergeneric hybrid of Gesneriaceae from Brazil. Phytotaxa 497:79–96. https://doi.org/10.11646/phytotaxa.497.2.2
Otto SP, Whitton J (2000) Polyploidy incidence and evolution. Annual Rev Genet 34:401–437. https://doi.org/10.1146/annurev.genet.34.1.401
Ramsey J, Schemske DW (1998) Pathways, mechanisms, and rates of polyploid formation in flowering plants. Annual Rev Ecol Systc 29:467–591. https://doi.org/10.1146/annurev.ecolsys.29.1.467
Renny-Byfield S, Wendel JF (2014) Doubling down on genomes polyploid and crop plants. Amer J Bot 101:1711–1725. https://doi.org/10.3732/ajb.1400119
Retallack GJ (2004) Late Miocene climate and life on land in Oregon within a context of Neogene global chenge. Palaeogeogr Palaeoclimatol Palaeoecol 214:97–123. https://doi.org/10.1016/j.palaeo.2004.07.024
Rieseberg LH (1997) Hybrid origins of plant species. Annual Rev Ecol Syst 28:359–389. https://doi.org/10.1146/annurev.ecolsys.28.1.359
Rieseberg LH, Ellstrand NC (1993) What can molecular and morphological markers tells us about plant hybridization? Crit Rev Pl Sci 12:213–241. https://doi.org/10.1080/07352689309701902
Rieseberg LH, Willis JH (2007) Plant speciation. Science, New Series 317(5840):910–914. https://doi.org/10.1126/science.1137729
Rieseberg LH, Archer MA, Wayne RK (1999) Transgressive segregation, adaptation, and speciation. Heredity 3:363–372. https://doi.org/10.1038/sj.hdy.6886170
Romero Zarco C (1986) A new method for estimating karyotype asymmetry. Taxon 35:526–530. https://doi.org/10.2307/1221906
Ronquist F, Huelsenbeck JP (2003) Mrbayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) Mrbayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Sáinz Ollero H, Moreno Saiz JC (2002) Flora vascular endémica española. In: Pineda FD, de Miguel JM, Casado MA, Montalvo J (coords) La Diversidad Biológica de España, Prentice Hall, Madrid, pp 175–195
Saito Y, Möller M, Kokubugata G, Katsuyama T, Marubashi W, Iwashina T (2006) Molecular evidence for repeated hybridization events involved in the origin of the genus Crepidiastrixeris (Asteraceae) using RAPDs and ITS data. Bot J Linn Soc 151:333–343. https://doi.org/10.1111/j.1095-8339.2006.00513.x
Sampaio GA (1913) Manual da flora portuguesa. Tipografia Occidental, Porto. Available at: http://bibdigital.rjb.csic.es/spa/Libro.php?Libro=24. Accessed 15 May 2016
Seehausen O (2013) Conditions when hybridization might predispose populations for adaptive radiation. J Evolution Biol 26:279–281. https://doi.org/10.1111/jeb.12026
Sehrish T, Symonds VV, Soltis DE, Soltis PS, Tate JA (2015) Cytonuclear coordination is not immediate upon allopolyploid formation in Tragopogon miscellus (Asteraceae) allopolyploids. PLoS ONE 10:1–17. https://doi.org/10.1371/journal.pone.0144339
Sharbrough J, Conover JL, Tate JA, Wendel JF, Sloan DB (2017) Cytonuclear responses to genome doubling. Amer J Bot 104:1277–1280. https://doi.org/10.3732/ajb.1700293
Sigel EM, Windham MD, Pryer KM (2014) Evidence for reciprocal origins in Polypodium hesperium (Polypodiaceae): a fern model system for investigating how multiple origins shape allopolypoid genomes. Amer J Bot 101:1476–1485. https://doi.org/10.3732/ajb.1400190
Siopa C, Dias MC, Castro M, Loureiro J, Castro S (2020) Is selfing a reproductive assurance promoting polyploid establishment? Reduced fitness, leaky self-incompatibility and lower inbreeding depression in neotetraploids. Amer J Bot 107:526–538. https://doi.org/10.1002/ajb2.1441
Smissen RD, Thorsen MJ, Breitwieser I, Ward JM (2015) DNA sequence analysis confirms the identity of the intergeneric hybrid Argyrotegium mackayi × Leucogenes leontopodium (Asteraceae:Gnaphalieae). New Zealand J Bot 53:210–221. https://doi.org/10.1080/0028825X.2015.1093000
Soejima A, Wen J, Zapata M, Dillon MO (2008) Phylogeny and putative hybridization in the subtribe Paranepheliinae (Liabeae, Asteraceae), implications for classification, biogeography, and Andean orogeny. J Syst Evol 46:375–390. https://doi.org/10.3724/SP.J.1002.2008.08065
Soltis DE, Soltis PS (1999) Polyploidy:recurrent formation and genome evolution. Tree 14:348–352. https://doi.org/10.1016/S0169-5347(99)01638-9
Soltis DE, Soltis PS, Tate JA (2004) Advances in the study of polyploidy since plant speciation. New Phytol 161:173–191. https://doi.org/10.1046/j.1469-8137.2003.00948.x
Soltis DE, Albert VA, Leebens-Mack J, Bell CD, Paterson AH, Zheng C, Sankoff D, dePamphilis CW, Wall PK, Soltis PS (2009) Polyploidy and Angiosperm diversification. Amer J Bot 96:336–348. https://doi.org/10.3732/ajb.0800079
Soltis DE, Visger CJ, Soltis PS (2014) The polyploidy revolution then…and now: Stebbins revisited. Amer J Bot 101:1057–1078. https://doi.org/10.3732/ajb.1400178
Soltis DE, Visger CJ, Marchant DB, Soltis PS (2016) Polyploidy: Pitfalls and paths to a paradigm. Amer J Bot 103:1146–1166. https://doi.org/10.3732/ajb.1500501
Spaniel S, Marhold K, Filová B, Zozomová-Lihová J (2011) Genetic and morphological variation in the diploid-polyploid Alyssum montanum in Central Europe: taxonomic and evolutionary considerations. Pl Syst Evol 294:1–25. https://doi.org/10.1007/s00606-011-0438-y
Stebbins GL (1938) Cytological characteristics associated with the different growth habits in the dicotyledons. Amer J Bot 25:189–198. https://doi.org/10.2307/2436589
Stebbins GL (1950) Variation and evolution in plants. Columbia University Press, New York. https://doi.org/10.7312/steb94536
Stebbins GL (1971) Chromosomal evolution in higher plants. Edward Arnold, London. Available at: https://wellcomecollection.org/works/pgk97y88
Stebbins GL (1985) Polyploidy, hybridization and the invasion of new habitats. Ann Missouri Bot Gard 72:824–832. Available at: https://wellcomecollection.org/works/pgk97y88
Struwe L, Kadereit JW, Klackenberg J, Nilsson S, Thiv M, von Hagen KB, Albert VA (2002) Systematics, character evolution, and biogeography of Gentianaceae, including a new tribal and subtribal classification. In: Struwe L, Albert VA (eds) Gentianaceae systematics and natural history. Cambridge University Press, Cambridge, pp 21–309
Stuessy TF (2009) Plant taxonomy: the systematic evaluation of comparative data, 2nd edn. Columbia University Press, New York
Suda J, Trávníček P (2006) Estimation of relative nuclear DNA content in dehydrated plant tissues by flow cytometry. Curr Protoc Cytom 38(Unit7):30. https://doi.org/10.1002/0471142956.cy0730s38.10.1002/0471142956.cy0730s38
Te Beest M, Le Roux JJ, Richardson DM, Brysting AK, Suda J, Kubešová M, Pyšek P (2012) The more the better? The role of polyploidy in facilitating plant invasions. Ann Bot (Oxford) 109:19–45. https://doi.org/10.1093/aob/mcr277
Thiers B (2016) [continuously updated] Index Herbariorum: a global directory of public herbaria and associated staff. New York Botanical Garden's Virtual Herbarium. Available at: http://sweetgum.nybg.org/ih/
Thompson JD (2005) Plant evolution in the Mediterranean. Oxford University Press, Oxford
Thompson JN, Nuismer SL, Merg K (2004) Plant polyploidy and the evolutionary ecology of plant/animal interactions. Biol J Linn Soc 82:511–519. https://doi.org/10.1111/j.1095-8312.2004.00338.x
Turner BL (1993) The Texas species of Centaurium (Gentianaceae). Phytologia 75:259–275. http://www.biodiversitylibrary.org/item/81266#page/77/mode/1up
Ubsdell RAE (1976). Studies on varition and evolution in Centaurium erythraea Rafn and C. littorale (D. Turner) Gilmour in the Britsih Isles 2. Cytology. Watsonia 11:7–31. https://archive.bsbi.org.uk/Wats11p33.pdf
Vallejo-Marín M, Buggs RJA, Cooley AM, Puzey JR (2015) Speciation by genome duplication: Repeated origins and genomic composition of the recently formed allopolyploid speciesMimulus peregrinus. Evolution 69:1487–1500. https://doi.org/10.1111/evo.12678
Vargas P, Carrió E, Guzmán B, Amat E, Güemes J (2009) A geographical pattern of Antirrhinum (Scrophulariaceae) speciation since the Pliocene based on plastid and nuclear DNA polymorphisms. J Biogeogr 36:1297–1312. https://doi.org/10.1111/j.1365-2699.2008.02059.x
Vilatersana R, Susanna A, García-Jacas N, Garnatje T (2000) Karyology, generic delineation and dysploidy in the genera Carduncellus, Carthamus and Phonus (Asteraceae). Bot J Linn Soc 134:425–438. https://doi.org/10.1111/j.1095-8339.2000.tb00539.x
Webb CJ, Druce AP (1984) A natural intergeneric hybrid, Aciphylla squarrosa Gingidia montana, and the frequency of hybrids among other New Zealand apioid Umbelliferae. New Zealand J Bot 22:403–411. https://doi.org/10.1080/0028825X.1984.10425272
Weiblen GD, Brehm BG (1996) Reproductive strategies and barriers to hybridization between Tellima grandiflora and Tolmiea menziesii (Saxifragaceae). Amer J Bot 83:910–918. https://doi.org/10.1002/j.1537-2197.1996.tb12784.x
Welles SR, Ellstrand (2016) Genetic structure reveals a history of multiple independent origins followed by admixture in the allopolyploid weed Salsola ryanii. Evol Appl 9:871–878. https://doi.org/10.1111/eva.12399
Wendel JF (2015) The wondrous cycles of polyploidy in plants. Amer J Bot 102:1753–1756. https://doi.org/10.3732/ajb.1500320
Whitney KD, Ahem JR, Campbell LG, Albert LP, King MA (2010) Patterns of hybridization in plants. Perspect Pl Ecol Evol Syst 12:175–182. https://doi.org/10.1016/j.ppees.2010.02.002
Widmer A, Lexer C, Cozzolino S (2009) Evolution of reproductive isolation in plants. Heredity 102:31–38. https://doi.org/10.1038/hdy.2008.69
Wood TE, Takebayashi N, Barker MS, Mayrose I, Greenspoon PB, Rieseberg LH, Crane PR (2009) The Frequency of polyploid speciation in vascular plants. Proc Natl Acad Sci USA 106:13875–13879. https://doi.org/10.1073/pnas.0811575106
Wu W, Zhou R, Huang Y, Boufford DE, Shi S (2010) Molecular evidence for natural intergeneric hybridization between Liquidambar and Altingia. J Pl Res 123:231–239. https://doi.org/10.1007/s10265-009-0275-z
Zeltner L (1970) Recherches de biosystèmatique sur les genres Blackstonia Huds. et Centaurium Hill (Gentianacées). Bull Soc Neuchâtel Sci Nat 93:1–164
Zeltner L (1991) Contribution to l’étude cytogographique des genres Blackstonia Huds. et Centaurium Hill. (Gentianaceae) en Turquie, à Rhodes et à Chypre. Bull Soc Neuchâtel Sci Nat 114:77–103
Zenil-Ferguson R, Ponciano JM, Bureigh G (2016) Evaluating the role of genome downsizing and size thresholds from genome size distributions in angiosperms. Amer J Bot 103:1175–1186. https://doi.org/10.3732/ajb.1500408
Acknowledgements
We thank Dra. C. Santa-Bárbara and Agronomist J. Díaz Fernández, who guided us in the field research, and L. Ligenfert for assisting in seed sowing. We also acknowledge the valuable taxonomic and phylogenetic advice of Prof. B. Valdés and Dr. J. Viruel. We are very grateful the financial support for the molecular analyses by a grant from the MINECO (Ministerio de Economía, Industria y Competitividad) research project (CGL2013-45037-P), led by Prof. J. Arroyo, and for the cytometry analyses by the research project CULTIVAR (CENTRO-01-0145-FEDER-000020), co-financed by Centro 2020, Portugal 2020 and EU through European Fund for Regional Development (ERDF). We appreciate the help of Herbarium SEV for preserving dry and frozen material and for consulting specimens from herbaria. We especially thank E. Folhadela curator of the Herbário Departamento de Botânica, Universidade do Porto (PO), for providing photographic images of the specimen type of G. Sampaio. This research has been possible thanks to the availability of digital images online of herbarium-type specimens, for which we greatly appreciate the public access to this information to herbaria BM, BR, E, GH, HAL, JE, K, NY and P, whose digitization projects facilitate taxonomic research.
Funding
Funding for open access publishing: Universidad de Sevilla/CBUA. Financial support was received for the molecular analyses by the Grant from the MINECO (Ministerio de Economía, Industria y Competitividad) research project (CGL2013-45037-P), and for the cytometry analyses by the project CULTIVAR (CENTRO-01-0145-FEDER-000020), co-financed by Centro 2020, Portugal 2020 and EU through European Fund for Regional Development (ERDF).
Author information
Authors and Affiliations
Contributions
ZDL framed the starting hypothesis and designed the experimental phase, taking part in the sampling and different analyses of every phase of the research, made the taxonomic revision, and wrote the first and the final draft integrating the contributions of the rest of authors. ME designed, developed and analysed the molecular analyses, contributing significantly to the taxonomic decision, so as to the writing and revision of the final draft. CAC put forward in the sampling and experiments in the field, in the seed germination experiments and contributed to the revision of the final draft. CG-L took part in the sampling and experiments in the field, the seed germination experiments, and in the morphologic and molecular analyses of the plants. JL and SC developed and edited the cytometry analyses and contributed significantly to writing and revision of the final draft. All of the authors read and approved the final manuscript of this article.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Handling Editor: Mike Thiv.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Specimens and accession number in GenBank for DNA sequences used in the molecular analyses.
Online Resource 2. Results for the PCA analysis: Eigenvalue, percentage and cumulative variance for the first 5 principal components, and eigenvectors for the continuous characters in them.
Online Resource 3. Results for the OSA analysis: Eigenvalue, percentage of variance for the two dimensions, and their eigenvectors for the nine categorical characters.
Online Resource 4. Results for the ANOVA analysis of the differences in 25 continuous characters among Exaculum pusillum, Schenkia elegans and S. spicata.
Online Resource 5. Mean and 95 % confidence interval for some of the characters showing significant differences at p<0.0001 in the ANOVA carried out to the population means of Exaculum pusillum, Schenkia elegans and S. spicata.
Online Resource 6. Maximum and minimum values of the chromosome length (in μm) for the largest and the lowest chromosome pairs in the karyogram, and values of the ratio (r) between both, following Stebbins (1971), per each population studied in Exaculum pusillum, Schenkia spicata and S. elegans. Range, mean and standard deviation (s.d.) of values are included for each taxon.
Online Resource 7. Phylogenetic tree of tribe Chironieae, inferred using ITS sequences.
Online Resource 8. Phylogenetic tree of tribe Chironieae, inferred using plastid sequences.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Díaz Lifante, Z., Escudero, M., Andrés Camacho, C. et al. Phylogenetic, cytogenetic and morphological evidences are critical for recognizing a new genus: Valdesiana, an Iberian intergeneric allopolyploid between Schenkia and Exaculum. Plant Syst Evol 309, 30 (2023). https://doi.org/10.1007/s00606-023-01864-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-023-01864-0